Are you a journalist? Please sign up here for our press releases
Subscribe to our monthly newsletter:
For the past two decades, life science researchers have been using libraries to borrow genes and proteins, rather than books. Some of the most frequented libraries are those containing series of yeast cells. Yeast are used in research labs around the world not only because they are required in the bread, wine and beer industries, but mostly because their cells are extremely similar to ours, making them excellent models for understanding human cell biology. Weizmann Institute of Science researchers have now developed a new system for creating yeast cell libraries. With this system, any lab can create a tailor-made library for almost any protein research.
A yeast library may contain thousands of “volumes” of genetically altered yeast strains. One library can have mutations in each of the 6,000 yeast proteins, while another may contain a collection of strains with fluorescent labels attached to their proteins. Such libraries have become the workhorse of modern yeast research, used to study the various functions of proteins or where they reside in the cell. But even though yeast cells contain many fewer genes than a human cell’s 25,000, making new libraries has been tedious and expensive, and until now, few have been created.
Our understanding of the cell has been something like having all of the parts of a car laid out on the garage floor
The Weizmann researchers, including Prof. Maya Schuldiner, Ido Yofe and Uri Weill of the Molecular Genetics Department, set out to change that situation. Ultimately, Weill, Yofe, Schuldiner and collaborators Prof. Michael Knop of the Zentrum für Molekulare Biologie der Universität Heidelberg (ZMBH) in Germany and his postdoc Dr. Anton Khmelinskii, together with the Weizmann Institute’s Dr. Emmanuel Levy and Dr. Ehud Sass, made a “mother” library that can be used to make any other library – rapidly, efficiently and cheaply. The approach is based on “swapping” the original tag to any other genetic piece in the yeast genome using an enzyme, so the team named this method SWAT (Swap Tag). This technique makes library construction something like playing with Lego – mixing and matching the volumes to put together ideal libraries. To showcase this approach, the team made seven brand new libraries for all of this organism’s proteins – nearly as many as had been made by all yeast labs in the past 20 years.
One of the things that Schuldiner and her team hope to accomplish with the new libraries is to gain a better understanding of the localization of protein in the cell. “Our understanding of the cell has been something like having all of the parts of a car laid out on the garage floor. We still needed the information that could tell us how to put them together to construct a moving car,” says Weill. “For that, we need to create the ‘diagram’ that shows us where the ‘axle’ fits in relation to the ‘steering wheel’ and the ‘body of the car’.”

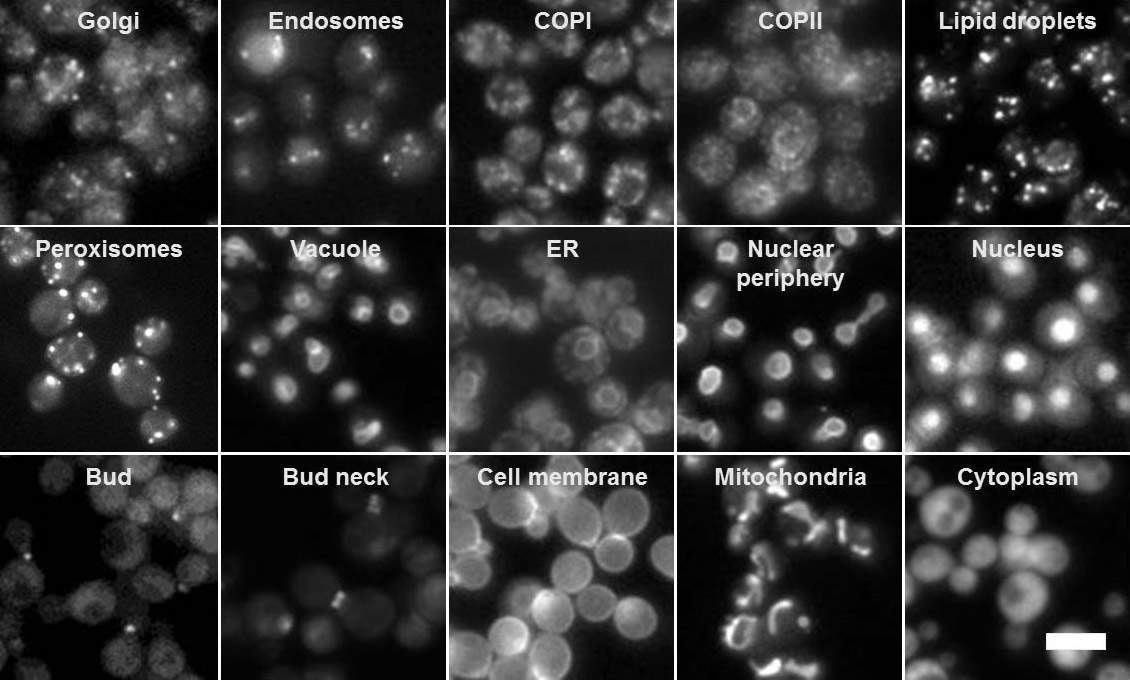
That lack was, in part, because in the existing library in which all the yeast proteins were labeled with a fluorescent protein, only around half of the proteins could be assigned a locale. Moving a fluorescent tag to another part of the protein, however, enabled the team to create a library in which all the proteins could be tracked around the yeast cells. Using this approach, Schuldiner and her lab team managed to finally find the placement of nearly 2000 proteins; none of these had been seen before. For example they found new proteins locating themselves within the cells’ power stations – mitochondria – and the team have begun to investigate what function these proteins serve there.
Weill then went on to make several additional, diverse libraries, including, for example, one in which all of the yeast proteins were overexpressed. And then he and the team made one in which a second colored fluorescent tag was inserted – a useful feature for experiments involving the actions of several proteins. Finally, they created two more libraries in which only half a fluorescent tag was inserted into each protein – t hat is, the label only glows when a protein bearing the other half makes contact. Scientists using these libraries can now take proteins from complementary libraries and produce them together in yeast cells to see if they interact and – just as importantly – where in the cell they go when they meet.

“We spent over three years developing these libraries,” says Weill. “We tagged the proteins and photographed the yeast cells with our robotic system, and then went over the images one by one. In addition to finding where proteins are situated – which is our lab’s focus – we have created a versatile tool that should enable anyone in the world to assemble their own collections. Rather than years of effort, researchers using our libraries will be able to set up and run experiments inside of a month.”
Indeed, many labs around the world have already begun using the new yeast protein libraries and techniques in their research. “With the seven new libraries, we have more than doubled the number of libraries – and their contents – available for research,” says Schuldiner. “We expect many new discoveries in our lab and others to arise out of these new yeast protein libraries and the methods we developed. It’s something like moving from a tiny, neighborhood library to the main branch and being allowed to explore all of the stacks at once.”
Prof. Maya Schuldiner's research is supported by the Edmond de Rothschild Foundations; the Foundation Adelis; and the Berlin Family Foundation Prof. Schuldiner is the incumbent of the Dr. Gil Omenn and Martha Darling Professorial Chair in Molecular Genetics.
Dr. Emmanuel Levy's research is supported by the David and Fela Shapell Family Foundation INCPM Fund for Preclinical Studies; the Richard Bar Laboratory; the Merle S. Cahn Foundation; and Anne-Marie Boucher and Mitch Garber. Dr. Levy is the incumbent of the Recanati Career Development Chair of Cancer Research in Perpetuity.