Are you a journalist? Please sign up here for our press releases
Subscribe to our monthly newsletter:
Genes, like people, form “social” networks: They interact with numerous other genes, which, in turn, interact with yet others. Such interactions form the basis for biological entities called biological pathways, which make up the life of each living cell. Learning more about biological pathways and their genes is therefore essential for understanding how the human body works, but obtaining information on the pathways can become extremely complex. The human genome has tens of thousands of genes, each of which may be connected with hundreds of others. Compounding the difficulty is the lack of a standardized approach: Just like human social networks, which partly overlap with one another and link people in different ways, various genetic databases provide information on pathways using different yardsticks for inclusion and different group naming systems.
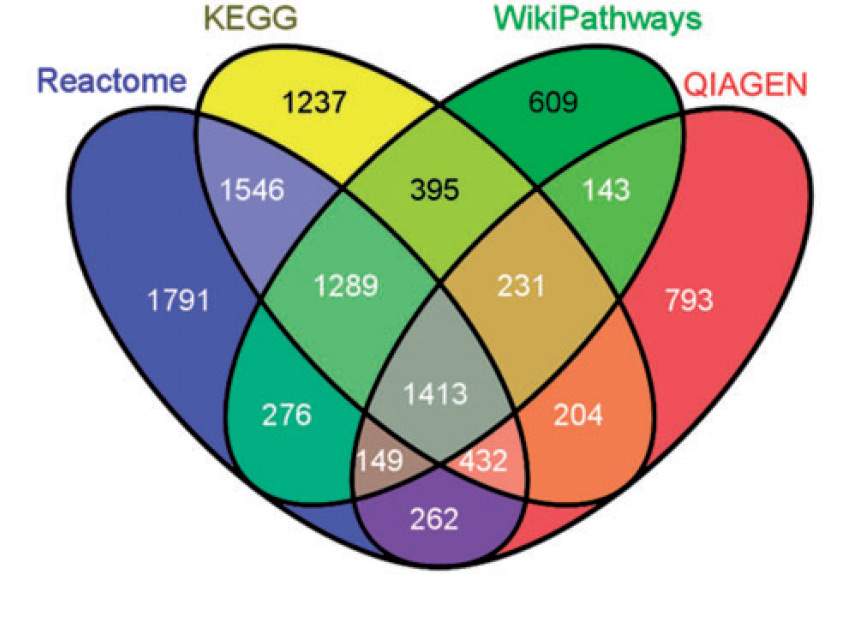
Weizmann Institute scientists, headed by Prof. Doron Lancet, have now created a universal resource of readily retrievable information on pathways and the genes they contain. Using a single, coherent mathematical approach that minimizes redundancy and maximizes information content, they integrated data on more than 3,000 pathways from a dozen bioinformatics resources into a single online database called PathCards. This global, consolidated repository provides a much more manageable number of unified pathways, about 1,000, referred to as SuperPaths. By optimizing the access to pathway information in this manner, PathCards can greatly facilitate basic and biomedical research.
As described recently in the journal Database (Oxford), PathCards supplies extensive connectivity data on more than 10,000 genes, building upon and complementing GeneCards, the database of all human genes created, nurtured and maintained by Lancet and his team at the Weizmann Institute for nearly two decades. The research was spearheaded by Dr. Frida Belinky, a postdoctoral fellow in the laboratory until recently. The authors also include Noam Nativ, Dr. Gil Stelzer, Shahar Zimmerman, Dr. Tsippi Iny Stein and GeneCards team head Marilyn Safran. The research is funded by the company LifeMap Sciences, which is headed by Dr. David Warshawsky.
By bringing numerous biological pathways effectively under one roof, PathCards, in a sense, presents the entire genome as a complex web of smaller “social networks.” And just as it is conceivable that two people who do not know each other may become associated through a friend who happens to belong to two disparate networks, genes get linked to one another in new ways through the joining of several pathways into a single SuperPath.

Using PathCards, scientists can perform quick, in-depth searches in all human SuperPaths. For example, a search for the insulin gene, the hormone that regulates sugar metabolism, provides a list of more than 160 relevant SuperPaths. But a researcher seeking a connection between insulin and, for example, adrenaline, will quickly uncover a subset of six SuperPaths that associate insulin with the adrenalin receptor, thus bringing the two hormones under one roof and displaying their mutual relationship in detail.
One of the ultimate goals of this project is to help scientists identify unsuspected genetic connections that make a difference – for example, discovering new disease genes that could constitute targets for developing novel drugs. PathCards can attain this goal by uncovering “guilt by association,” whereby if gene A is already known to be associated with a disease, one can infer that gene B, which shares a pathway with it, is a likely new candidate for a related biomedical function. Lancet and his team have also developed another bioinformatics tool, VarElect, that can be used in conjunction with GeneCards and PathCards to help discover culprit genes when sequencing the genomes of diseased individuals.
Prof. Doron Lancet’s research is supported by the Crown Human Genome Center, which he heads; the Nella and Leon Benoziyo Center for Neurological Diseases; the Wolfson Family Charitable Trust; the Dr. Dvora and Haim Teitelbaum Endowment Fund; the estate of Rosa and Ernst Guttmann; and LifeMap Sciences. Prof. Lancet is the incumbent of the Ralph D. and Lois R. Silver Professorial Chair of Human Genomics.