Are you a journalist? Please sign up here for our press releases
Subscribe to our monthly newsletter:
Proteins are the workhorses of the cell, carrying out the vast majority of the tasks required to keep us alive and healthy. But if they are damaged, proteins can form toxic clumps in the brain that are called amyloid fibers, which have been tied to Alzheimer’s, Parkinson’s and numerous other neurodegenerative diseases. Until recently, it was thought that once amyloids form, there is no going back. It turns out, however, that cooperation between two families of proteins can actually dissolve amyloids, allowing cells to return to a healthy state.

In two recent, back-to-back papers, Dr. Rina Rosenzweig from the Structural Biology Department at the Weizmann Institute of Science, together with researchers from Heidelberg University (ZMBH) and the German Cancer Research Centre (DKFZ), discovered the mechanism by which these two protein families – called Hsp70 and Hsp40 chaperones – break the amyloids apart. This action fits in with the chaperones’ general mission in the cell, which is to keep other proteins in line.
In the first paper, Dr. Anne Wentink, from Prof. Bernd Bukau’s group at ZMBH, found that the key to dissolving amyloid fibers was achieving a dense accumulation of Hsp70 chaperones in one spot. Once there, the Hsp70 molecules latch onto individual proteins in the amyloid and pull at them until they are freed. In the other paper, Drs. Ofrah Faust and Meital Abayev-Avraham from Rosenzweig’s group discovered a unique regulatory mechanism that enables the second chaperone family, Hsp40, to generate the clusters of Hsp70s required for breaking up the amyloid fibers.
Until recently, it was thought that once amyloids form, there is no going back
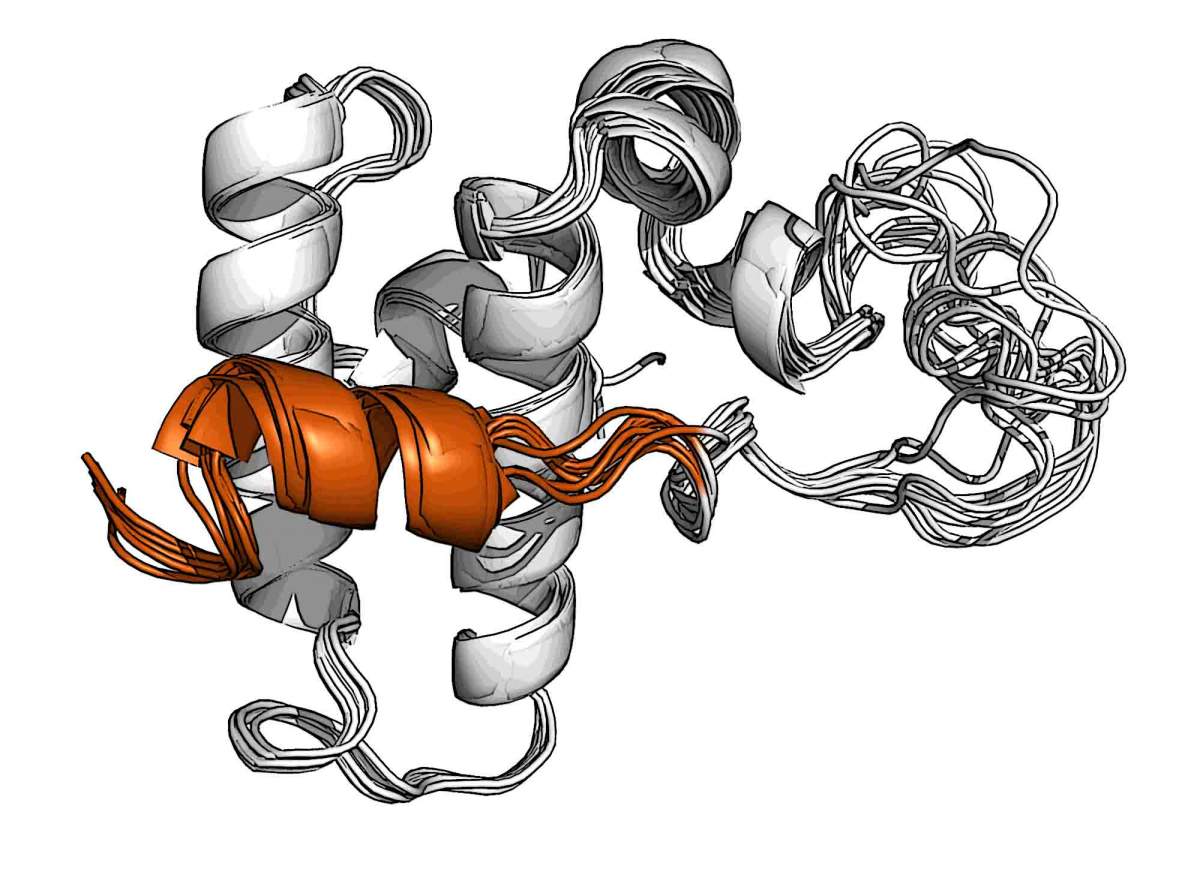
Using cutting-edge nuclear magnetic resonance spectroscopy methodologies and the advanced instrumentation in the Weizmann Institute’s Clore Institute for High-Field Magnetic Resonance Imaging and Spectroscopy, the Weizmann scientists were able to obtain the structure of one specific member of the Hsp40 family, called DnaJB1. This chaperone binds to the amyloid fibers and assists Hsp70 in dissolving them. An analysis of its structure revealed that DnaJB1 is uniquely capable of performing this task because it acts as a two-step switch of a previously unknown kind.
The scientists found that DnaJB1 contains a unique helical structure that temporarily blocks the binding site for Hsp70 chaperones. When Hsp70 molecules approach, they at first attach to DnaJB1 with their disordered region, which the researchers called a “tail.” This attaching action releases the helical block, enabling Hsp70 to connect to its proper binding site on DnaJB1, where it become activated and engages in pulling the amyloid fibers apart. DnaJB1 then releases the Hsp70 and reactivates the lock for the next chanperone, freeing itself to snare other Hsp70s and activate them on the same spot on the amyloid fiber. This repeated action generates a cluster of the chaperones large enough to dissolve the amyloids.
The discovery of these molecular details raises the possibility that progression of neurodegenerative diseases may be tied, among other things, to the disruption of this unique two-step regulatory mechanism – and this, in turn, opens a previously unexplored avenue of research in the battle against neurodegeneration. Moreover, the study’s findings may shed new light on other vital cellular processes, as the scientists predict that similar active helical structures are likely to exist in additional Hsp40 chaperones.
Participants in this study included members of Rosenzweig’s group in the Structural Biology Department at the Weizmann Institute, members of Bernd Bukau’s group at the University of Heidelberg, and Dr. Nir London, of Weizmann’s Organic Chemistry Department.
Dr. Rina Rosenzweig's research is supported by the Abisch Frenkel Foundation for the Promotion of Life Sciences; the Blythe Brenden-Mann New Scientist Fund; and the European Research Council.