Are you a journalist? Please sign up here for our press releases
Subscribe to our monthly newsletter:
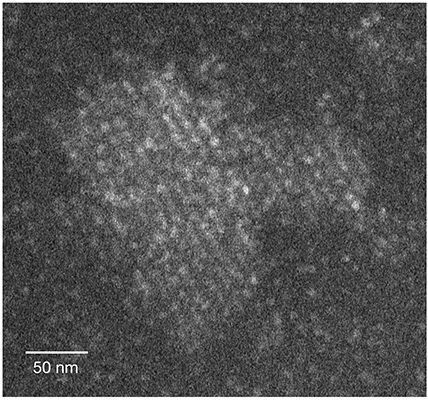

When we say that something is “crystal-clear,” we mean that it couldn’t possibly be clearer, but much about crystals themselves is not clear at all. Scientists are for the most part still unable to predict what kind of crystal a given molecule can produce, if at all; even the so-called “classical” theory of crystal formation is currently being questioned. In a study reported recently in Nature, researchers at the Weizmann Institute of Science have for the first time directly observed the crystallization process at the molecular level in three dimensions – and revealed a mechanism that helps explain crystallization cases that don’t fit the classical model. Because crystals are everywhere – from bones to mollusk shells in nature, and from pharmaceuticals to TV displays among man-made structures – knowing how they form is essential: It can help us understand how order emerges from disorder in the living world and advance the design of materials for a variety of uses.
In fact, it’s hardly surprising that crystallization is so difficult to figure out. After all, the process is practically counterintuitive: In nature, just as in our closets, creating order usually takes effort. So how does the orderly structure of crystals come about, for example, when they form spontaneously in a solution? The classical theory of crystallization, proposed in the early 20th century, holds that when molecules randomly bump into one another in a solution, they sometimes bond, their energetically beneficial state causing them to become a tiny crystalline nucleus that gradually grows into a larger, ordered crystal. But in the past thirty years or so, with the advancement of microscopy and other imaging methods, scientists around the world began to see instances of crystallization that didn’t fit into the classic model. In some cases, they observed crystals that started out as an amorphous aggregate, which suggested that the classical theory had to be revised – and raised questions as to what really happens when crystals form.
Prof. Boris Rybtchinski of the Organic Chemistry Department and his team have in the past few years gained unique expertise in obtaining direct images of crystal formation. In the new study, they set themselves the challenging goal of observing the crystallization of proteins – extremely large, complex organic macromolecules.

“We wanted to make observations without preconceived ideas – without trying to support this or that theory – and the optimal way to do that was to follow, in three dimensions, the location of every single molecule throughout the process,” Rybtchinski explains. “Though it’s technically impossible at present to visualize each one of most organic molecules as they undergo crystallization, very large proteins can be tracked down.”
It’s possible that there’s a spectrum of crystallization options in nature – from the classic model to the nonclassical process we observed with ferritin
Drs. Lothar Houben and Sharon G. Wolf of Weizmann’s Chemical Research Support Department suggested resorting to cryoSTEM tomography, a technique previously developed by them and Prof. Michael Elbaum of Chemical and Biological Physics Department at the Weizmann Institute of Science. “Cryo” stands for “cryogenic,” meaning that the sample is instantaneously frozen (vitrified) prior to the imaging; it is then scanned from different angles by an electron beam of a scanning transmission electron microscope, or STEM. The approach, followed by a mathematical analysis of the findings, enabled Houben – together with Wolf and Dr. Haim Weissman of the Organic Chemistry Department – to reconstruct the spatial location of each protein molecule in the sample. The protein in question was ferritin, which stores iron in various body tissues. After using cryo-STEM tomography to scan samples of a vitrified solution of ferritin at different stages throughout its crystallization and determining each molecule’s position, the scientists were able to assess the distances between the molecules, their density in the sample and how much their arrangement differed from an ideally formed crystal.
They found that ferritin crystallized in a decidedly nonclassical manner. Rather than immediately forming a perfect crystalline nucleus, it created an aggregate characterized by a very minor degree of order, which gradually released the water molecules it contained, bringing protein molecules closer to one another and increasing progressively in density and in the degree of order. The aggregate was initially more ordered in the center than on the borders, but with time, density and order grew on the periphery as well. In other words, order evolved gradually, spreading outward from the core.
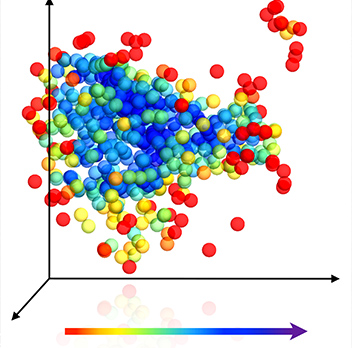
These findings suggest that the classical theory of crystallization is not necessarily wrong, but that it explains only a certain number of cases, in which a small crystalline nucleus does form right away. In other cases, however, the processes starts with a relatively disordered aggregate that progresses toward an ordered crystal.
“It’s possible that there’s a spectrum of crystallization options in nature – from the classic model to the nonclassical process we observed with ferritin,” Rybtchinski says. “It’s also possible that crystallization is always gradual but can proceed at different speeds, so that when order is created quickly, it seems to occur in an instant, as suggested by the classical model.”
Protein crystallization is crucial to the production of protein-based drugs such as insulin, and to solving protein structure by X-ray crystallography; crystallization in general is important in the pharmaceutical industry, as well as in materials science and in numerous other fields. The study’s findings may one day help bring crystallization under control, thus streamlining industrial processes.
Prof. Boris Rybtchinski is Head of the Solo Dwek and Maurizio Dwek Research School of Chemical Science. His research is supported by the Henri Gutwirth Fund for Research; and Dana and Yossie Hollander.