Are you a journalist? Please sign up here for our press releases
Subscribe to our monthly newsletter:

Can one ever tell what’s in a pitcher just by looking at its shape? In the field of structural proteomics – the study of protein structure – shape is the key to understanding a protein’s biological activities. “Solving the spatial structure of proteins gives scientists information on protein functions and properties that’s impossible to obtain in any other way. With this information, we can raise new scientific questions, design advanced experiments and point the way to more efficient drug design,” says Prof. Joel L. Sussman, Head of the Israel Structural Proteomics Center (ISPC) at the Weizmann Institute.
Indeed, working in close collaboration with research scientists to advance the study of proteins and their functions in the body has been the hallmark of the ISPC since its inception five years ago. This approach, which pairs the protein-solving expertise of the ISPC team with scientific groups investigating the actions of specific proteins, was ahead of its time; it has since been adopted by a number of such proteomics centers around the world. The Israeli center, founded by Institute Profs. Sussman, Gideon Schreiber, Israel Silman and Yigal Burstein, is a central node in a European network of structural proteomics research centers. With support from Israel’s Ministry of Science, the Divadol Foundation and the EU, it offers its services to research scientists at the Weizmann Institute and other research institutes in Israel and around the globe, as well as to physicians and researchers in the biotechnology and biomedical industries.
“Thanks to the work of the ISPC team,” says Schreiber, “the Weizmann Institute is well in the lead of the enormous worldwide effort to unravel the mysteries of protein structure.” This team, including Drs. Orly Dym, Tamar Unger, Shira Albeck and Yoav Peleg, sees each project through from beginning to end – from laying out the initial research plan through the various stages of cloning the gene encoding the protein, isolating and purifying it, crystallizing it, determining the structure of the crystallized protein and, finally, creating a 3-D model of the protein structure. What makes the work challenging is that each protein has a unique “personality,” and creating just the right conditions to get it to turn into a crystal and reveal that personality often takes ingenuity and patience.
In addition to solving protein structures, the ISPC offers assistance and advice to students and research groups, and also produces a number of proteins for use in biochemical and biomedical research. Of the more than 100 proteins purified by the team in the past five years, and the 40 protein structures they have solved, several have recently appeared in various scientific publications:
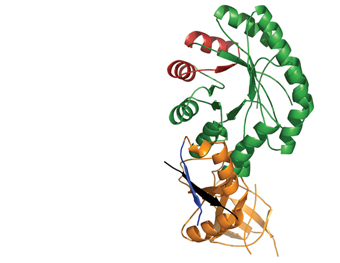
Polyamines are small molecules found in all living cells; they’re necessary for such fundamental cellular processes as the production of proteins and nucleic acids, fixing chromosome structure and regulating gene expression. Too low a level of polyamines inhibits cellular proliferation, while overly high levels can have a toxic effect. Cells contain a network of proteins whose concerted action keeps polyamine levels optimal. In a study that appeared in Protein Science, the ISPC team, together with Prof. Chaim Kahana of the Molecular Genetics Department, revealed details of the structure and working mechanisms of a key regulator that sets the rate for polyamine production.
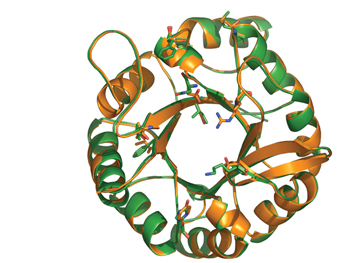
Prof. Alexander Bershadsky of the Molecular Cell Biology Department and his team research the activities of a protein, p120, that is involved in creating contact points between neighboring cells, and which appears to stimulate the formation of protrusions that enhance cell motility. The ISPC team succeeded in purifying p120 along with another protein, cortactin, involved in the assembly of the actin fibers that play a role in forming the protrusions. In research that was published in the Proceedings of the National Academy of Sciences (PNAS), USA, Bershadsky’s group used the purified proteins to show that p120 directly regulates cortactin activity, thus elucidating the role of this protein in cell motility.
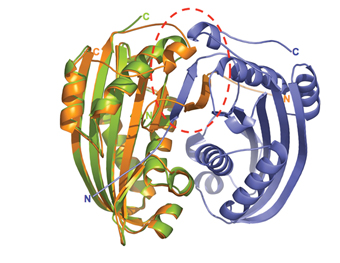
The bacterium Bacillus thuringinesis kills insects by destroying their cell membranes. In a collaborative effort, the ISPC team, together with Prof. Arieh Zaritsky and Shmulik Cohen of Ben-Gurion University of the Negev, solved the 3-D structure of one of the bacterial proteins responsible for its lethal action. These findings suggested to them the poison’s mode of action, as well as an explanation for it high toxicity. The study, which appeared in the Journal of Molecular Biology, may help researchers develop new natural pesticides, as well as drugs to destroy cancer cells.