
Real life is always more complicated than a model. Take, for example, the structure of a protein: Chemical modifications alter its basic shape, and then it is changed by interactions with other molecules, and even by the viscosity and acidity of surrounding fluids. These structural changes, in turn, affect a protein’s function, so studying its structure in real-life conditions – that is, inside a living cell – is essential for an in-depth understanding of numerous biological processes.
Until now, however, existing methods have only been suited to studying protein structure in artificial conditions – after the proteins are removed from the cell and crystallized or placed in a solution. Inside cells, scientists can sometimes determine protein structure using nuclear magnetic resonance. But this technology is unsuitable for exploring large proteins or solving the structure of proteins at their regular concentration in the cell; enhancing this concentration to see them better, again, creates conditions that are far removed from real life.
Now Weizmann Institute scientists have developed
a method for investigating protein structures in conditions that are close to real life. As reported in the
Journal of the American Chemical Society,
Prof. Daniella Goldfarb of the Weizmann Institute’s Chemical Physics Department and her team – Dr. Andrea Martorana and Dr. Akiva Feintuch from her own group and Dr. Giuliano Bellapadrona from the group of Prof. Michael Elbaum of the Materials and Interfaces Department, together with colleagues in Italy – have successfully applied this method to examining the structure of a common protein inside the cell.
The scientists employed a type of spectroscopy known as double electron-electron resonance, or DEER, which in the past had been used extensively to explore protein structure outside of cells, in frozen solutions. It works by measuring the distance between two locations in the protein. These two points are each labeled with a marker called a spin label, which has an odd number of electrons and therefore responds to the application of a magnetic field in combination with microwave irradiation. The challenge was that the most common such marker, nitroxide, gains another electron once inside the cell, thus becoming invisible to the magnetic field.
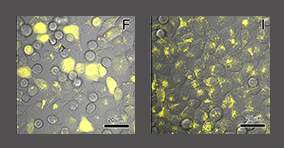
Building on their vast experience with spectroscopy in general and DEER in particular, Goldfarb and her team made use of a new, different marker: a charged atom of gadolinium, which commonly serves as a contrast agent in magnetic resonance imaging in medical applications and is known to be stable in the body. Such probes do not become “invisible” in the cell, but they pose a challenge of a different sort: They emit strong signals only in a high magnetic field, 10 times higher than that used in standard DEER experiments. Goldfarb had in the past developed such a high-field spectrometer for other protein studies, and now the scientists have adapted it to DEER measurements for the study of protein structure. Using gadolinium as a spin label, they first labeled a common protein called ubiquitin outside the cell and then delivered it back into the cell. They managed to investigate its structure in great detail inside frozen human cells, at a resolution of a few angstroms (ten-billionths of a meter). Importantly, ubiquitin’s concentration inside the cell was close to its levels in real life. The cells were not damaged by the freezing; in fact, after being thawed, they regained their full function.
Studying its structure in real-life conditions – that is, inside a living cell – is essential for an in-depth understanding of numerous biological processes.
Ubiquitin, whose job in the cell is to mark unwanted proteins for destruction, is such an essential molecule that its structure has long been solved outside the cell, but the new method now furnishes biologists with the ability to observe changes in its structure in real-life conditions.
Now that the new method has been shown to work for ubiquitin, it can be applied to other proteins. It may prove particularly valuable in the many biological processes in which proteins alter their shape and folding, so that their real-life structure at various stages of the process are quite different from that solved outside the cell. Such structural alterations occur, for example, when proteins transport drug molecules across the cellular membrane, participate in the metabolism of calcium and other ions, or form the abnormal plaques responsible for Alzheimer’s disease.
Prof. Daniella Goldfarb’s research is supported by the Carolito Stiftung; and the De Benedetti Foundation-Cherasco 1547. Prof. Goldfarb is the incumbent of the Erich Klieger Professorial Chair in Chemical Physics.

