Are you a journalist? Please sign up here for our press releases
Subscribe to our monthly newsletter:
The very name “disordered proteins” suggests that something is off about protein molecules that hang loose spaghetti-style instead of being neatly folded into 3D structures. But a new study at the Weizmann Institute of Science, reported in the Proceedings of the National Academy of Sciences (PNAS), shows that such proteins are good at adapting to changes in the cell’s environment and continue to function well despite their unstructured state.
“Disordered proteins are found much more often in higher organisms than in bacteria,” says team leader Dr. Hagen Hofmann of the Structural Biology Department. “This relatively late enrichment in the course of evolution suggests that they signal a shift toward greater complexity and sophistication, but why exactly does nature need them?”
When scientists started discovering disordered portions of protein molecules in the late 1980s and early 1990s, they would chop them off and discard these bits, as they couldn’t be crystalized for structural study. With time, however, scientists found that the disordered state is much more prevalent than they had believed: It applies to about 30% of the entire set of proteins in an organism, affecting parts or even the entire length of the molecule in many proteins. Moreover, it turned out that disordered proteins form an important class of biological molecules that are thought to take part in controlling gene activity and in a host of other vital functions. In fact, they may even have advantages over ordered ones in certain situations. They are easier to break down as needed or to be modified in a changing environment, and they are more likely to have multiple functions because their floppy structure is literally more open to binding with other molecules than the “traditional” 3D architecture that requires a precise fit.
Disorder applies to about 30% of the entire set of proteins in an organism
In other words, disordered proteins are so flexible in structure and function; but are they also sufficiently sturdy to perform important tasks? In the new study, Dr. Renee Vancraenenbroeck and Yair Harel from Hofmann’s lab, in collaboration with Prof. Wenwei Zheng from Arizona State University, addressed this question by testing whether disordered proteins are sensitive to changes in the cell’s environment. Focusing on a network of more than a hundred proteins that control gene expression in the cell, they selected five disordered proteins that work together, playing key roles within this network. The scientists attached red and green fluorescent probes to both ends of each protein’s molecule, submerged the molecules in a solution and viewed them under a microscope while changing the salt concentration in this solution, to mimic the fluctuations in salt concentration that commonly occur within living cells. The ratio of green to red fluorescence told them whether the molecules underwent structural changes.
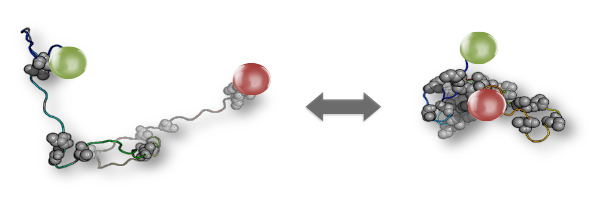
The researchers found that all the five disordered proteins were sensitive to the changing environment: Their 3D structure changed with the fluctuations in the solution. But surprisingly, theoretical calculations performed on the experimental data revealed that these changes had no effect on the functioning of the protein network. This happened because the five proteins underwent the changes in synchrony with one another – for example, they all expanded or shrank in response to a particular change – so that the ultimate performance of the network remained the same.
“We’ve shown that disordered proteins are sensitive to their environment,” Hofmann says. “But we’ve also shown that the network they form compensates for this sensitivity and remains sufficiently stable to continue functioning despite the changes.”

These findings suggest that disordered proteins form sturdy networks that are well equipped to perform a variety of vital tasks in the cell. And this, in turn, can lead to a better understanding of cellular processes in which these proteins play an important role. For example, among highly disordered proteins are those encoded by several genes that play central roles in cancer, including the master tumor suppressor p53 and the oncogene gene c-Myc.
Dr. Hagen Hofmann's research is supported by the Leir Foundation; and the European Research Council. Dr Hofmann is the incumbent of the Corinne S. Koshland Career Development Chair in Perpetuity.