Are you a journalist? Please sign up here for our press releases
Subscribe to our monthly newsletter:
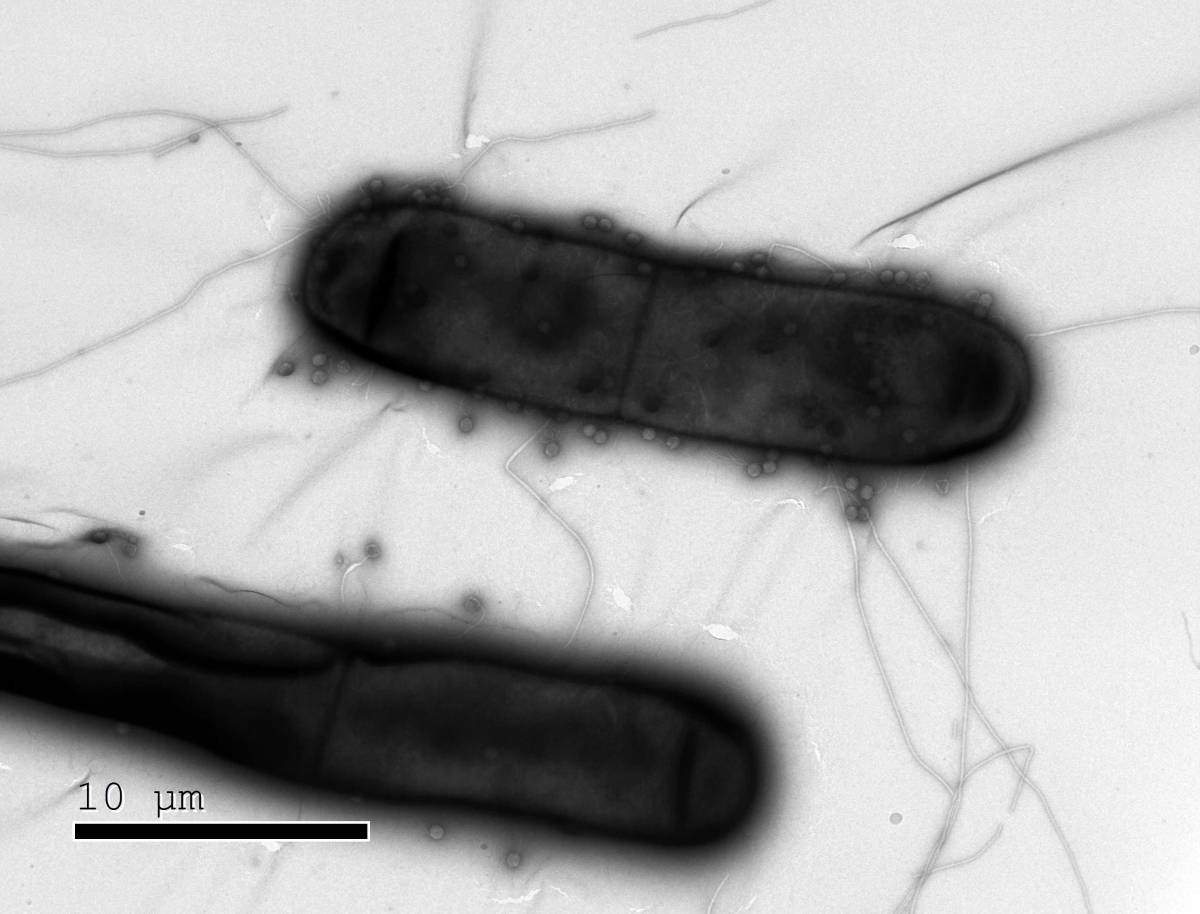
Until a decade ago, scientists were not aware that bacteria had complex immune systems – ones that could keep up with the pace of evolution in viruses called phages that infect bacteria. That changed with the discovery of what is now the most famous bacterial immune mechanism: CRISPR. Scientists realized that CRISPR is a natural gene editor, and it has revolutionized the world of biological research in thousands of labs around the world. Researchers now understand that most microorganisms have sophisticated immune systems of which CRISPR is just one element; but there has been no good way to identify these systems. In a massive, systematic study, Prof. Rotem Sorek and his team at the Weizmann Institute of Science have now revealed the existence of 10 previously unknown immune defense mechanisms in bacteria. “The systems we discovered are unlike anything we had seen before,” says Sorek. “But among them, we think, are one or two that might have the potential to increase the gene-editing toolbox, and others that point to the origins of the human immune system.” The results of their study were recently published in Science.
Bacteria cannot rely on CRISPR alone in the war against phages, explains Sorek, a member of the Institute’s Molecular Genetics Department. Indeed many phages have “anti-CRISPR” proteins that cancel CRISPR activity, suggesting that other systems take up the slack. Sorek and his team began their search for these systems by creating a computer program that would scan all the bacterial genomes that have ever been sequenced – around 50,000 genomes in all. Rather than look for sequences with predefined characteristics, the algorithms they created searched for the “statistical signatures” of genes involved in defense – for example, their location in “defense islands” where several defense-related genes are found near one another. Then, because immune system genes rarely work alone – even in bacteria – the researchers developed complex computer analytic methods so as to understand which genes join forces and work together to form a defense system.
One or two might have the potential to increase the gene-editing toolbox
Once they had narrowed down the possible defense genes from millions to several hundred, the researchers needed to test the candidate mechanisms they had identified. Rather than attempting to isolate the genetic sequences from hundreds of different bacteria, the team turned to synthetic biology: getting the genes made to order. They sent the strings of gene code – totaling something like 400,000 bases, or “letters” of genetic code – to a commercial lab where dozens of different multi-gene systems were synthesized for testing. These synthetic systems were inserted into lab bacteria that had had their natural immune systems inactivated. The bacteria were then exposed to phages and other infective elements to see if the transplanted defense system was a viable one. Out of the various systems the researchers examined, 10 strongly protected the lab bacteria from infection, thus identifying them as new immune defense systems.
Sorek says that between the various stages of computer analysis and the experimentation, the study required the intensive effort of six people working for two years in his lab. The study was led by Dr. Shany Doron and Sarah Melamed, with the intensive involvement of Gal Ofir, Dr. Azita Leavitt, Dr. Anna Lopatina and Dr. Gil Amitai. The team took to meeting every other week for a “defense council” to discuss the different arms of the research and the defense mechanisms they had discovered.
Others point to the origins of the human immune system
The researchers still don’t know how the new bacterial immune systems function, and some according to Sorek, “seem to have surprising functions that we are now starting to investigate.” One of these systems contains Toll-Interleukin Receptor (or TIR) domains. TIR domains were already known to be involved in immune systems – but until now not that of microorganisms. These domains are an integral part of the human immune system and even that of plants, but they had never before been shown in bacteria to be involved in anti-viral defense. “Our findings show that some of the important parts of our own immune system have deep evolutionary roots in the bacterial immunity mechanisms,” says Sorek.
Other genes seemed to have been “borrowed” from non-defensive bacterial systems. One of them, for example, is known from the flagella that bacteria use to swim. These genes provide the flagella with energy by enabling them to take in protons; one of the new defense systems found in Sorek’s lab uses these genes it its protection against phages. Another one, called condensin, generally protects the DNA during cell division, and the researchers discovered a defense system that uses components of the condensin mechanism to protect bacteria against the invasion of plasmids – tiny rings of DNA that can parasitize the bacterial cells.

“The fact that we managed to find 10 new bacterial defense systems implies there are even more out there,” says Sorek. “My lab is continuing to search for new ones. In addition, we are starting to focus on several of the more promising ones to understand how they function.”
Sorek says that the new discoveries are exciting because of the new windows they provide on the evolution of immune systems and the eternal battle between viruses and the organisms they infect. But he also believes that some may turn out to be powerful tools for biological research: “Every immune system, by definition, needs to target invading elements in a very specific yet flexible way, and we can use this targeting for biotechnological purposes – as we’ve done with CRISPR and with restriction enzymes before it. Any one of the new systems we found might be the next gene-editing tool – or perhaps even the foundation of even more exciting molecular tools,” says Sorek.
Prof. Rotem Sorek's research is supported by the David and Fela Shapell Family Foundation INCPM Fund for Preclinical Studies; the Y. Leon Benoziyo Institute for Molecular Medicine; the Dana and Yossie Hollander Center for Structural Proteomics; the Abisch Frenkel Foundation for the Promotion of Life Sciences; the Leona M. and Harry B. Helmsley Charitable Trust; Martin Kushner Schnur; and the European Research Council.