About 3 million people die every year as a result of infection with HIV, the virus that causes AIDS. Efforts to develop a vaccine against the virus are up against a particularly clever enemy: HIV assumes different guises even after entering the human body, going through rapid genetic changes that allow it to escape the immune system. In fact, HIV undergoes more changes while residing in the body of a single person than the flu virus has undergone since it was first discovered.
HIV-1, the type of HIV responsible for 90% of all infections, comes in two principal forms, called R5 and X4. These differ from each other significantly, both in the type of immune cells they attack and in the progression of the disease. R5 binds to proteins on the surface of the immune cells known as macrophages, weakening the immune system in the latent stage of the disease, when the person is a carrier of the virus but has not yet developed full-blown AIDS. By contrast, X4 attaches to the membranes of different immune system cells called T cells, causing these cells to die rapidly and bringing on the fatal stage of the syndrome, in which the immune system collapses completely. Yet for all the functional disparity between these two versions of the virus, when it comes to their constitutional sequence of amino acids – the building blocks of proteins – they differ in only one or two amino acids found in a certain protein on the viral envelope.
How does a change in a single amino acid affect the virus’s attack mechanisms to such an extent? In a study reported in the Proceedings of the National Academy of Sciences (PNAS), Prof. Jacob Anglister and research students Osnat Rosen and Michal Sharon of the Weizmann Institute’s Structural Biology Department used nuclear magnetic resonance (NMR) to closely examine the two versions of the virus. They focused on that segment of the envelope protein called V3 which is involved in the protein’s binding to target cell membranes – a critical stage in viral infection.
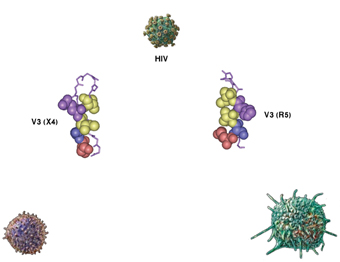
The scientists synthesized the sequence of amino acids making up the V3 protein segment in both versions and examined the structure produced when they bind to two different antibodies. They reasoned that in binding to an antibody, V3 would assume its natural spatial shape – the one that exists in the free virus and allows it to bind to various human cells. At first glance, the experiment seemed to produce two identical V3 structures, both shaped like a hairpin. However, a closer examination revealed a difference: Whereas the line-up of amino acids on one side of the hairpin was identical in both versions, on the other side, the orientation of X4 differed from that of R5. Consequently, the position of certain bonds between the amino acids on the two sides of the hairpin also changed. In addition, in X4, the amino acid side chains on one of the hairpin’s sides were pointed in the opposite direction to those of R5, resulting in significant differences in the protein’s shape and in the spatial organization of its binding site. Because of these variations in the shape of one crucial protein, the different versions of the virus latch on to different cells – macrophages in the case of R5, T cells in the case of X4 – and attack the immune system in different ways.
But what is it about the changes in the chemical sequence of the viral envelope protein that brings about the structural changes? And what makes this change so dramatic? To answer these questions, the scientists created a more complete picture of the spatial structure of V3, including the amino acid responsible for the differences between the two types of HIV. The picture revealed that this amino acid has a negative electric charge in R5, which allows it to interact with a positively charged amino acid on the opposite side of the hairpin, thereby stabilizing this structure. In contrast, a mutation in the X4 version causes the negatively charged amino acid to be replaced with a positively charged one, resulting in repulsion between the two parts of the hairpin: Here, the amino acids’ side chains, which carry the electric charge, turn on their axes and distance themselves from each other. Another mutation that can result in the transition of the virus from the R5 to the X4 form creates a sequence of three positively charged amino acids, which interacts with the electron clouds of another set of amino acids located diagonally across on the other side of the hairpin. The resulting structure is stable but different from the one characterizing the viral envelope protein in R5.
These findings reveal how a single mutation causes the virus to progress rapidly from one version of HIV to another. Moreover, they explain how mutations taking place in the virus inside the human body significantly affect the development of the disease and allow the virus to branch out in its attack strategy.
Prof. Jacob Anglister’s research is supported by the Joseph and Ceil Mazer Center for Structural Biology; Mr. Samy Cohn, Rio de Janeiro, Brazil; and Mr. Joe Gurwin, New York, NY. Prof. Anglister is the incumbent of the Joseph and Ruth Owades Professorial Chair in Chemistry.