Are you a journalist? Please sign up here for our press releases
Subscribe to our monthly newsletter:
“Anything found to be true of Escherichia coli must also be true of elephants,” said Jacques Monod, who had won the Nobel Prize in 1965 for describing how bacteria, such as E. coli, regulate their gene expression. Assuming Monod was right, why is the way in which genes are organized in bacteria so different from that of elephants and other eukaryotes – organisms whose cells contain a nucleus and other membrane-bound organelles?
Bacteria use a “beads on a string” approach to gene expression: Genes that encode for proteins participating in the same cellular process are organized one after the other along the bacterial DNA strand – similarly to a rosary – in a functional unit called an “operon.” In response to specific stimuli, these genes are then expressed together, as one long messenger RNA molecule. “The bacterial operon is a great way to ensure that the right genes are expressed together at the right time,” says Prof. Jeffrey Gerst of the Weizmann Institute of Science’s Molecular Genetics Department.

Unlike bacteria, eukaryotes – from yeast to humans – have evolved to store their genetic information in the nucleus (which prokaryotes, such as bacteria, lack by definition), where they keep their genes on different chromosomes, often with no apparent order or logic. This raises the question as to how the eukaryotic cell is able to coordinate the expression of several genes that are needed for the execution of a biological process. “Compared to the relatively straightforward bacterial arrangement, the eukaryotic gene organization is somewhat chaotic, so there has to be an explanation, an underlying mechanism that allowed complex life forms, like elephants, to evolve over time and thrive,” adds Gerst.
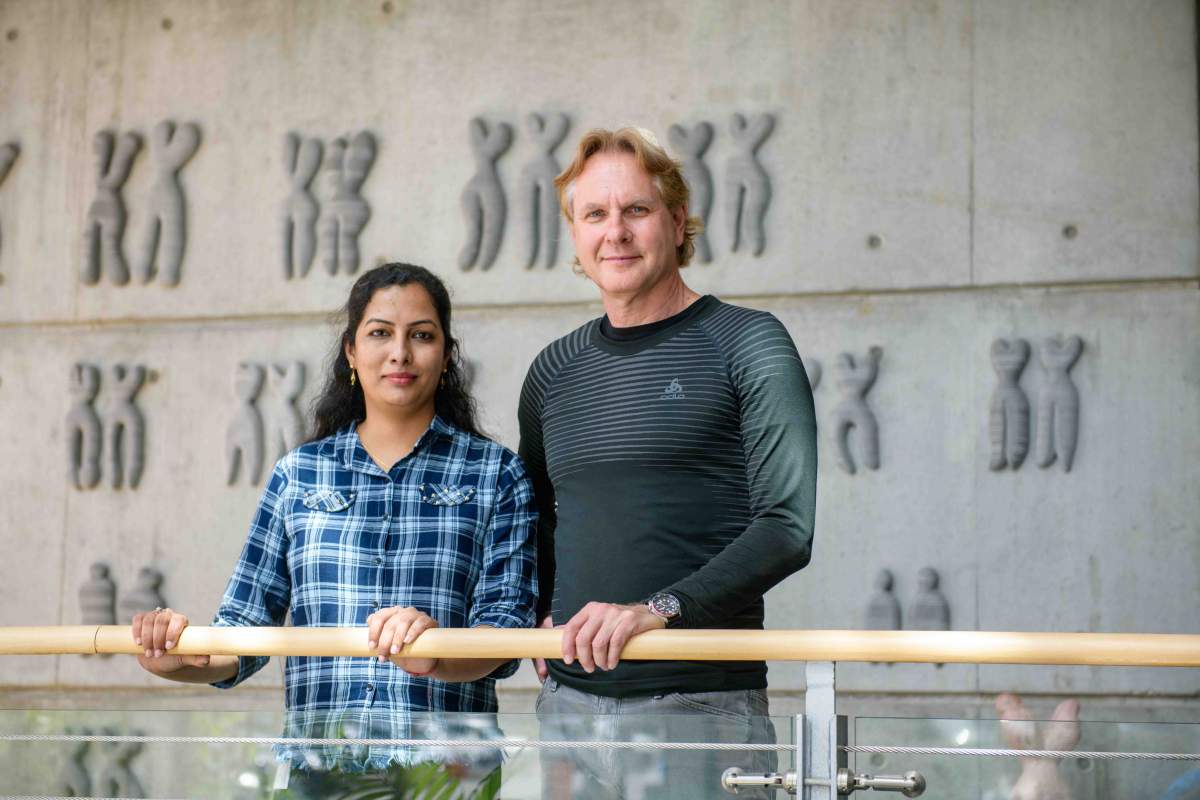
To address this question, Gerst and his team, spearheaded by postdoctoral fellow Dr. Rohini Nair and graduate student Dmitry Zabezhinsky, turned to baker’s yeast. Known by its Latin name Saccharomyces cerevisiae, this single-cell eukaryote has been used by humankind for many years – not only to make delicious food and intoxicating beverages but also as a model organism for studying processes that are believed to have been conserved throughout evolution. The research team used an approach previously developed by Gerst and colleagues to capture messenger RNAs by precipitating, or “pulling-down,” specific proteins that bind to the RNA molecules upon leaving the nucleus. In this way the researchers were, for the first time, able to track down and analyze a bundle of RNA molecules that originate from different chromosomes but participate in the same biological process in yeast.
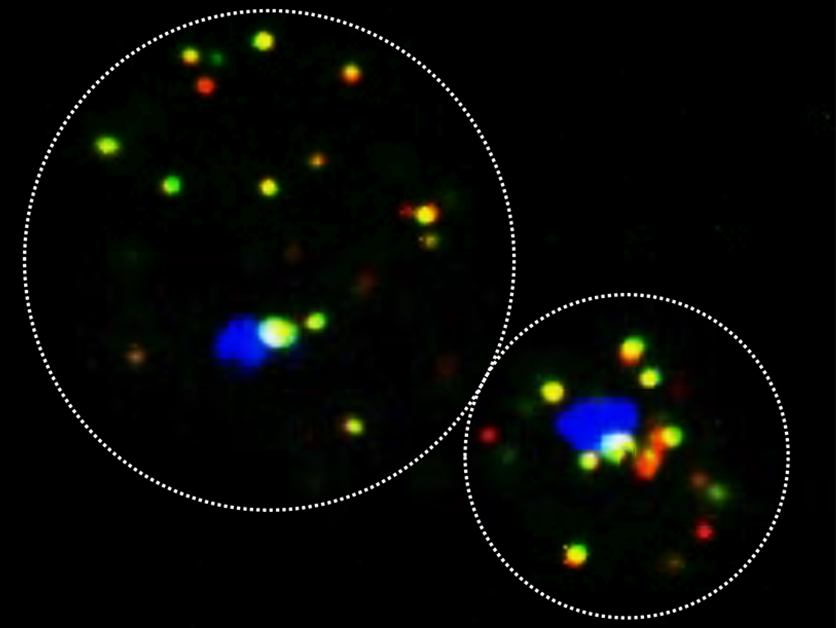
“We named these RNA bundles “transperons,” to describe messenger RNAs that have originated from different chromosomes but act as a sort of functional unit, like the bacterial operon,” says Gerst. Nair then tested the newly formulated “transperon” hypothesis on a universal cellular process known as the “heatshock response,” which is performed by a group of different proteins in response to unfavorably warm temperatures. The team then found that for this cellular process, the messenger RNAs appear to have assembled together as a “transperon” bundle. “These findings point to a possible early evolutionary event that must have granted eukaryotes the flexibility needed for creating new cellular processes,” says Gerst.
""Compared to the relatively straightforward bacterial arrangement, the eukaryotic gene organization is somewhat chaotic, so there has to be an explanation, an underlying mechanism that allowed complex life forms to evolve"
The study has also found that in order to create transperons, yeast cells adjust the structure of their DNA to physically bring the genes needed for the same process closer together. In this way, the cells ensure that these genes will be expressed at more or less the same time and will end up together in the same bundle. “When we perturbed the proteins that help to assemble transperons, the RNA molecules disconnected from the bundle and the cellular process they were intended for was disrupted,” explains Gerst.
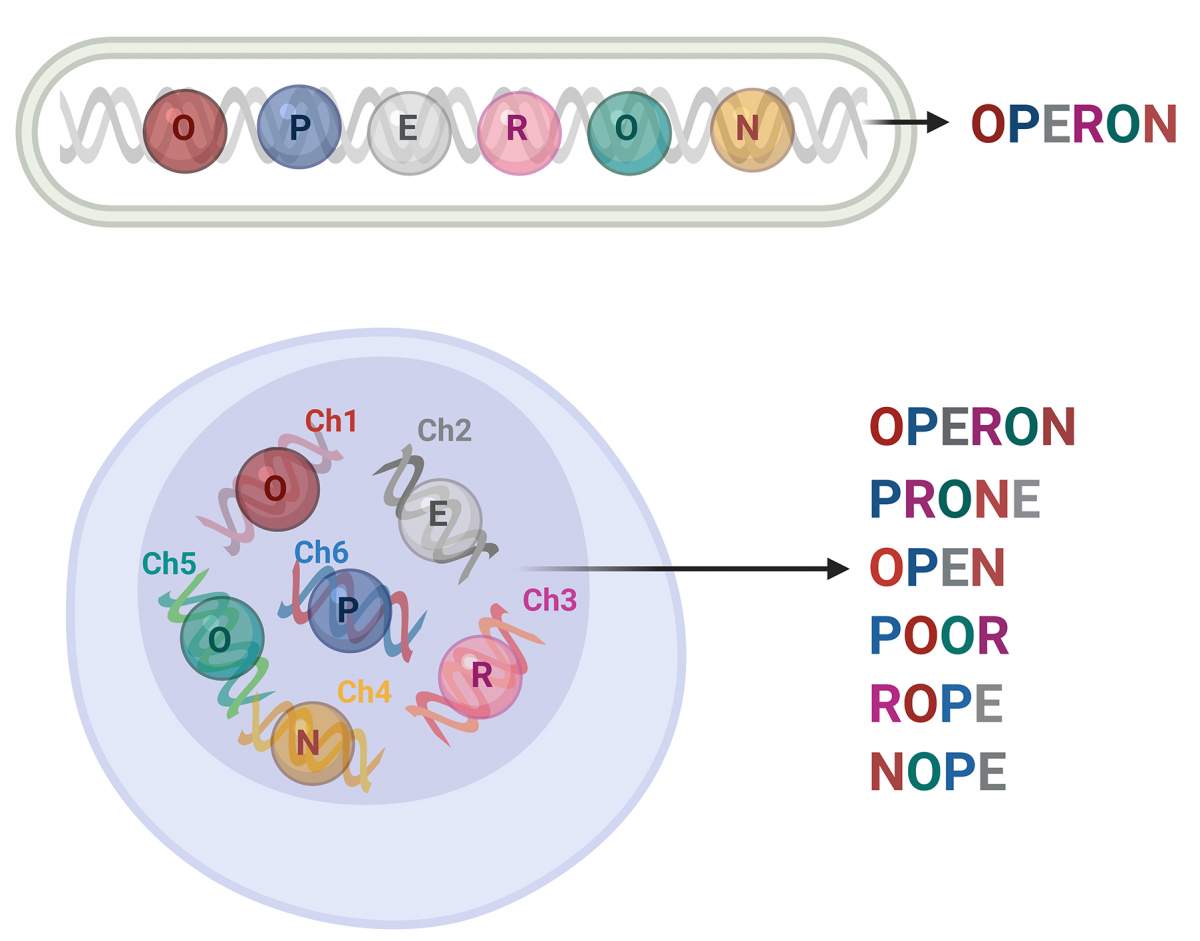
These findings suggest that yeast cells, and possibly other eukaryotes, have developed a form of genetic Scrabble, wherein they use the same “letters” – that is, RNA molecules expressed by genes located on different chromosomes – to form different “words” or “transperons.” Thus, eukaryotes have been able to expand their functional repertoire and evolve into complex life forms – from fungi and exotic plants to peacocks – but still maintain a degree of control and regulation over cellular processes like their bacterial ancestors. While Monod’s claim may have been exaggerated, elephants do seem, after all, to have a mechanism similar to that of bacteria for ensuring that things run smoothly in their cells – even if they use this mechanism in a different manner.
An average yeast cell, measuring 5 microns, contains 30,000 messenger RNA molecules, 200,000 ribosomes and more than 108 proteins.
Prof.Gerst’s research is supported by the Miles and Kelly Nadal and Family Laboratory for Research in Molecular Genetics.
Prof. Gerst is the incumbent of the Besen-Brender Professorial Chair of Microbiology and Parasitology.