The DNA strands in our genomes can pair up comfortably in double-helix fashion, but what about the lonely, single-stranded RNA that is copied from the DNA? The “letters” in the RNA sequence, nucleotides, are chemically built like those in the DNA to seek a mate – sometimes even with another letter on the strand – causing the strand to curl and bend over on itself into irregular hairpin-shaped configurations.
A new study performed by Weizmann Institute and Stanford scientists and published in
Nature suggests that these shapes seem to act as a sort of “notation” on top of the gene code, helping the cell’s machinery to read out that code.
In possibly one of the broadest studies done on RNA structure in human cells,
Prof. Eran Segal and research student Ohad Manor in the Weizmann Institute’s Computer Science and Applied Mathematics, and Molecular Cell Biology Departments, joined up with the research team of Prof. Howard Chang of Stanford University to map the total set of RNA configurations in one type of cell, in three individuals. Using a technique Segal and his group had developed and patented in 2010, they created a scorecard for each sequence of RNA to locate the points where the segments double up and where they remain unattached. Analyzing over 160 million RNA fragments from each individual – all together thousands of RNAs – enabled them to a produce sort of panoramic, topographical map of the RNA landscape.
“Some things immediately stood out when we looked at the pairing scores,” says Segal. “For example, we can see very clearly the sharp dips at the ‘start’ and ‘stop’ points of the gene that demarcate its functional region.” Those dips – unpaired regions in the folded strand – are the hairpin-like curves and bulges in the structure. This nearly braille-like notation, says Segal, may be a handy way for such machinery as the ribosome, which produces proteins according to the RNA sequence, to find its place on the strand without having to perform a search for the right group of letters. In fact, the code hidden within the shapes may extend to the readouts for individual amino acids, each of which is encoded in a three-letter sequence known as a codon. The score showed a pattern of extra-close pairing every three letters that may signal the beginning of a new codon.
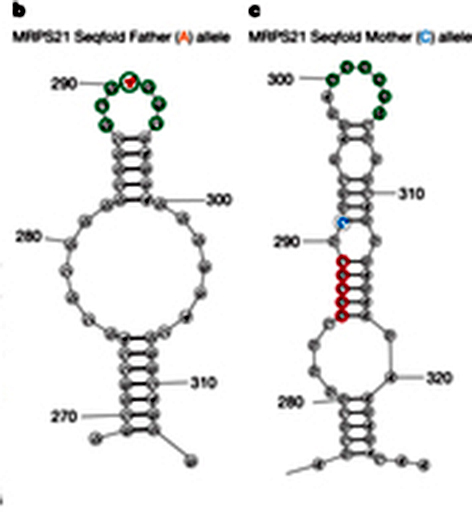
The three individuals were parents and child, a setup that enabled the researchers to ask some questions about the RNA landscape and heredity. They looked at short segments of the RNA where alterations in one or two letters are known to occur frequently, so the child would likely have inherited two different versions. What difference does a single letter substitution make? Comparing the pairing scores from mother, father and child showed that the shapes of around 15% of the RNAs containing these sequences were significantly altered – enough to substantially affect the function of the protein. The team named these configurations RiboSNitches.
The team then flagged the RiboSNitches they identified on the RNA map. The sites of these suggested to them that such RiboSNitches could play a crucial role in regulating the process in which the gene code is translated into protein. Their positions on the map even hint that some RiboSNitches may be involved in such diseases as multiple sclerosis, asthma and Parkinson’s.
Prof. Eran Segal’s research is supported by the Kahn Family Research Center for Systems Biology of the Human Cell; the Carolito Stiftung; the Cecil and Hilda Lewis Charitable Trust; and the European Research Council.