Are you a journalist? Please sign up here for our press releases
Subscribe to our monthly newsletter:
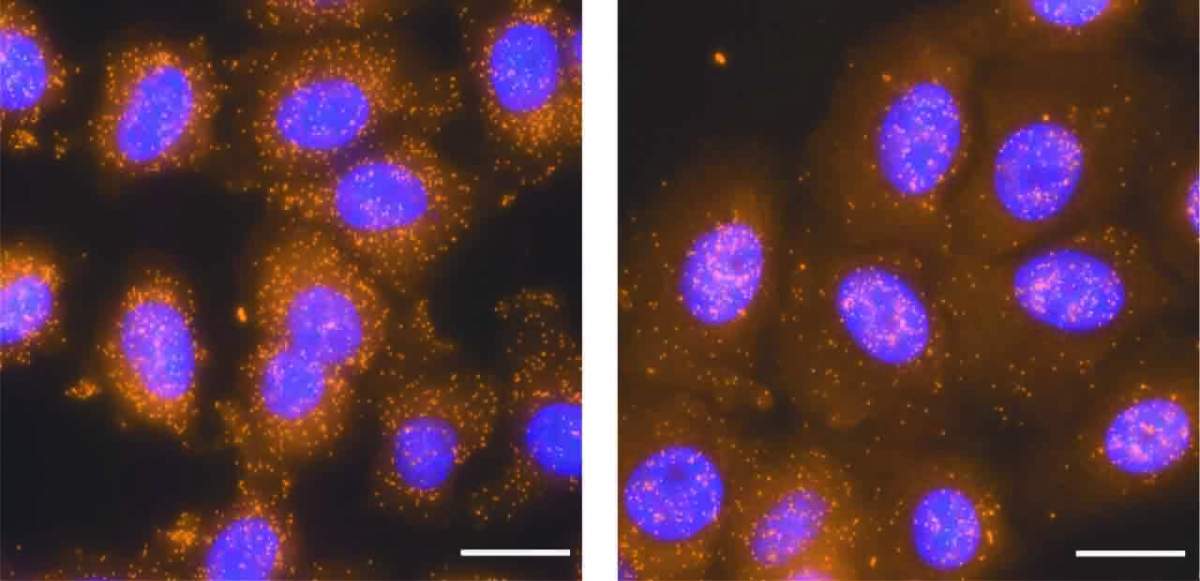
The way in which RNA is exported from the cell nucleus – a process that is crucial for all cellular life – was mostly thought to be uniform and fast. But in a new study, researchers at the Weizmann Institute of Science have shown that the cell uses at least two different mechanisms of RNA export; and the study explained how the export of particular RNAs may be delayed while others can still rapidly exit the nucleus.
The researchers also showed that the existence of the two mechanisms can allow viruses to perform a selective shutdown, blocking the part of the cell’s RNA machinery that’s responsible for antiviral defenses without harming the rest of the RNA, so as, potentially, to avoid killing the cell that serves as their home. These findings shed new light on fundamental genetic processes and may enhance our understanding of viral infection.
“Textbooks state that the export of virtually all RNAs relies on the same set of genes that make several key proteins, but we found that in human cells, there are at least two separate export routes, which gives both our cells – and viruses – the ability to shut off RNA export in a selective manner,” says team leader Dr. Igor Ulitsky of Biological Regulation Department. He conducted the study with graduate students Binyamin Zuckerman and Maya Ron, in collaboration with Dr. Martin Mikl and Prof. Eran Segal of the Molecular Cell Biology and Computer Science and Applied Mathematics Departments.
Viruses have apparently evolved to benefit from the existence of the two export pathways
Production of RNA in the cell nucleus is relatively slow, but its export – in which RNA is packaged and escorted from the nucleus to the cytoplasm – is normally so smooth that it takes just several minutes. In fact, there’s usually five times more RNA in the cytoplasm than in the nucleus, and it’s in the cytoplasm that most RNA serves as a template for manufacturing proteins. In the past few years, however, scientists found that sometimes, RNA gets stuck for quite some time within the nucleus, where it can perform functions that are unrelated to protein manufacture.
To explore this phenomenon, Ulitsky and colleagues selectively silenced several genes responsible for the export of RNA from the nuclei of human cells. They identified two groups of genes responsible for protein machineries that play different roles in this export, distinguishing between RNAs made up of few segments called exons, and RNAs that are spliced together from numerous exons. The number of exons in an RNA molecule can range from one to over one hundred. The scientists found that one of the two machineries, NXF1, exported mostly RNAs containing no more than three exons; the other, TREX, focused on RNAs containing four exons or more. When the researchers silenced NXF1, the few-exon RNAs got stuck in the nucleus; when they silenced TREX, a different group of RNA molecules got stuck.

The process of splicing several exons together probably developed in the course of evolution to help cells distinguish between their own RNA – most of which contains numerous exons – and that belonging to viruses or other foreign invaders, which typically has only one exon. The existence of the two different export routes for RNAs with different numbers of exons thus may provide the cell with flexibility in regulating RNA activity. The researchers found that in order to be escorted out of the nucleus, the few-exon RNAs had to contain a well-folded genetic segment that served as a zip code of sorts, potentially marking them as different from foreign or erroneously transcribed RNA strands and facilitating their move to the cytoplasm. Any few-exon RNA lacking the added fold would get stuck in the nucleus, unable to manufacture proteins that could endanger the cell. In contrast, since the multiple-exon RNA usually belongs to the cell itself, not to the foreign invaders, it gets a green light for being exported to the cytoplasm without needing this extra zip code, and is therefore less likely to get stuck within the nucleus.
Viruses have apparently evolved to benefit from the existence of the two export pathways. Upon infecting the cell, they can selectively prevent the export of few-exon RNAs – just the type of RNA that makes interferons, the first line of defense against viruses – but not the export of multiple-exon RNA. When the Weizmann researchers analyzed data from human cells exposed to viral proteins that block RNA export, they found that it was mainly the few-exon RNAs that got stuck in the nucleus, not the multiple-exon RNAs. This finding suggests that by performing a selective RNA shutdown, viruses prevent the cell from making interferons intended to fight the infection.
Dr. Igor Ulitsky's research is supported by the Sagol Institute for Longevity Research; the Willner Family Center for Vascular Biology; the Kekst Family Institute for Medical Genetics; the Abisch Frenkel Foundation for the Promotion of Life Sciences; the Minna-James Heineman Stiftung; the Blavatnik Award; and the European Research Council. Dr. Ulitsky is the incumbent of the Sygnet Career Development Chair for Bioinformatics.