Are you a journalist? Please sign up here for our press releases
Subscribe to our monthly newsletter:
How do the fingers and vertebrae in a growing embryo develop, or the stripes on a zebra’s hide arrange themselves? For over half a century, developmental scientists have used a model first created by Alan Turing that explains how complex biological patterns can emerge based on just a few simple building blocks. In Turing’s model, each cell in the growing embryo “knows” what to do next by sensing the other cells around it. New research at the Weizmann Institute of Science and the Università degli Studi di Firenze suggests that making Turing’s elegant model somewhat “noisier” can provide a picture of developmental pattern formation that is considerably more robust than the original. Their findings add to a growing body of evidence that, rather than disrupting biological development, such noise is a basic ingredient on which these systems are built.
Prof. Joel Stavans of the Weizmann Institute’s Physics of Complex Systems Department needed an experimental model system that would allow him to tackle basic questions of developmental pattern formation: “In an ideal system, one can interrogate each cell at every stage; it can be described by a small number of parameters and displays a high degree of control. Human embryos were out the question, since pattern formation in these is based on hundreds of cell types. Even the fruit flies found in many developmental biology labs have around 60 cell types. We needed something much more manageable. Fortunately, we found the system we sought in multicellular cyanobacterial microorganisms.”
Cyanobacteria are ubiquitous microorganisms that, long before plants arose, were churning out oxygen into the atmosphere and fixing the nitrogen that plants and animals require. More importantly, to Stavans and his team, they have only two basic cell types, and these grow in simple filaments.
The particular cyanobacterial organism that Stavans and his colleagues investigated, together with Prof. Duccio Fanelli and his group in Italy, represents one of the first experiments in multicellularity on Earth. In this organism’s filaments, cells are aligned one after the other. These cells are photosynthetic: They use the Sun’s light to produce oxygen and store energy in the form of carbohydrates. These cells also take up nitrogen to build proteins, and for this they prefer nitrogen that is in a readily usable form. But if the easy-to-use sources are depleted, cyanobacterial cells can “fix” the abundant nitrogen in the atmosphere.

They do this using a specialized enzyme, but this enzyme’s function is impaired by the oxygen that the cells produce in the process of photosynthesis. To cope with this inner contradiction, cells divide their labor: Some of the cells in the filament become so-called heterocysts that specialize in fixing atmospheric nitrogen, while the others continue to release oxygen through photosynthesis. Thus a developmental pattern arises in which single heterocysts are separated by intervals of about 10-15 photosynthetic cells -- one of the simplest patterns in nature.
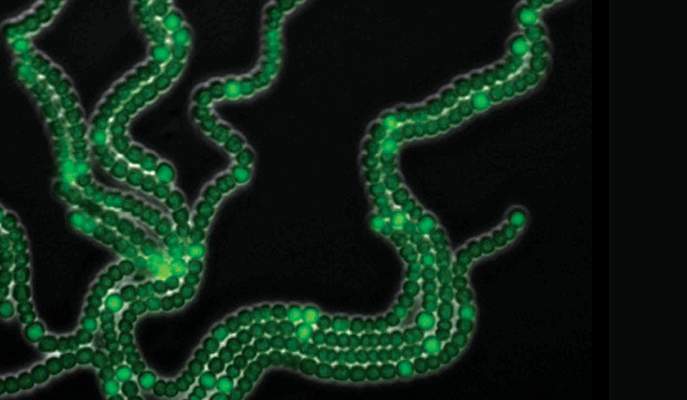
In their experiments, the researchers tagged genes involved in the differentiation process with added genes for fluorescent proteins. They then exposed cyanobacterial filaments to the nitrogen-poor conditions that produce the patterned filaments and observed the ensuing process in time-lapse movies. As the filaments grew, the cells that differentiated into heterocysts lit up under the microscope.
But how well did the observed development fit Turing’s neat conceptual scheme?
We are seeing that noise is a necessary condition, on top of the well-defined blueprint
“Turing’s model, as he conceived it, is fully deterministic. It yields patterns, but only over a very narrow range of parameter values; therefore the model requires fine-tuning to arrive at most developmental patterns. But even the patterns in the simplest multicellular cyanobacteria appear over a wide range of conditions, and the cells’ behavior we observed is far from deterministic. It is actually very noisy, and even neighboring cells in the filaments exhibited large differences in gene expression. So we added this feature to the Turing model – adjusting it for stochastic (noisy) fluctuations,” says Stavans. “The new version of the model produced a much a more robust description than the original, and it did not require that finicky ‘fine tuning’.”
The research group recently reported on their “stochastic Turing model” in PLoS Biology.
Taken together with other recent work, this study suggests that fluctuations and noise are at the very heart of many of the patterns we often think of as rigidly predetermined. They are built into the system from day one. In terms of the physics, Stavans explains, “living systems never reach equilibrium; they have to expend energy to enable accuracy in replicating information from one generation to the next. In biological systems we are seeing that noise is a necessary condition, on top of the well-defined blueprint, for them to grow and develop – to end up with quite precise shapes, arrangements and patterns.”
Prof. Joel Stavans is the incument of the Siegfried and Irma Ullmann Professorial Chair.