Enzymes are shape-shifters – their intricate molecular structures take on a variety of different arrangements as they go about their work. The ability to trace in real time the dynamic changes an enzyme molecule's structure undergoes has been a holy grail of molecular biology – one that scientists thought would only be attained many years down the road. But reality has overtaken the predictions: Prof. Irit Sagi of the Structural Biology Department in the Chemistry Faculty and her research team have developed a method of recording an enzyme's movements down to the scale of single atoms, and this technique is now being used as a tool for designing new drugs.
The complex molecular machines known as enzymes are involved in nearly all the functions of our bodies. Changes in enzyme structure take place at dizzying speeds – tiny fractions of a second – and this dynamic action makes them extremely efficient. To understand how enzymes work, scientists generally use a variety of techniques, such as crystallization, to determine the three-dimensional structure of the resting molecule. Although much information is revealed in these studies, they often can't give scientists a satisfactory picture of the steps involved in shifting an enzyme's shape. Yet a precise understanding of each step in enzyme activity can be especially important to drug makers, who aim to create new drugs that can tightly focus on a target protein or prevent a single type of action from taking place.
The Weizmann team's method allows them to identify the movement of single atoms residing within the active site of an enzyme molecule. The scientists freeze the process at various stages and then apply cutting-edge techniques borrowed from the field of X-ray spectroscopy and structural chemical analysis to determine the configuration of the molecule at each of those stages
.
One of the enzymes tested by the team's method is an enzyme called TNF alpha convertase, which is suspected of involvement in many diseases, from multiple sclerosis to cancer. TNF alpha convertase is a protease – part of an enzyme family that cleaves other proteins and prepares them for specific actions. Among other functions, TNF alpha convertase cleaves a protein called PRO-TNF alpha, slicing through the part of its structure that attaches it to the inside of the cell wall. The released PRO-TNF is then free to act in the cell. These proteins perform some crucial functions, but if too many are released at once, problems can start to occur and diseases can develop. For this reason, scientists in labs around the world have pinpointed TNF alpha convertase as a target for drugs that will damp down PRO-TNF activity.
The only problem is that drugs that block TNF alpha convertase activity tend to have disastrous, sometimes even fatal, side effects. The reason for this is the familial resemblance between the structure of this enzyme and that of other protease family members. It turns out that these drugs are not overly discriminating about which protease they block, and they end up hindering some vital functions in the cell. Identifying the real-time structural and biophysical events taking place during enzyme action could therefore be a real boon to drug research.
At this point, Sagi and her team, together with Dr. Marcos Milla of Roche Palo Alto, LLC, in California, entered the picture. With the dynamic observation method they had developed, they succeeded in recording every change in the molecule at intervals of a few thousandths of a second. In the first stage of their research, they were able to put an end to a scientific debate that had gone on for years by pinpointing a specific response mechanism the enzyme employs.
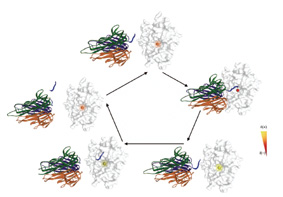
Next, they discovered that when the enzyme closes in on the protein that it's preparing to cleave, it begins to get "excited." As it makes contact with the surface of the protein, the dynamic structural transformations pick up their pace. This is especially evident in the electrons belonging to the lone zinc atom sitting squarely in the active site of the enzyme molecule. The changes they observed appear to be uniquely characteristic of TNF alpha convertase; if drugs could be designed to target this particular action, they might avoid affecting other proteases and thus prevent unwanted side effects. The results of this study appeared recently in the Proceedings of the National Academy of Sciences (PNAS), USA.
This technique might be applied to a great many of the enzymes that are involved in various disease processes. The ability to observe enzyme activities in fine detail may turn out to be a powerful tool that could lead to new approaches in drug design and new drugs that are more efficient and less likely to cause side effects. Various pharmaceutical companies have already expressed interest in the Institute team's new technique.
Prof. Irit Sagi's research is supported by the Helen and Milton A. Kimmelman Center for Biomolecular Structure and Assembly; the Avron-Wilstaetter Minerva Center; Mr. and Mrs. Michael Ambach, Boca Raton, FL; and the estate of David Turner. Prof. Sagi is the incumbent of the Maurizio Pontecorvo Professorial Chair.