Are you a journalist? Please sign up here for our press releases
Subscribe to our monthly newsletter:

"Jack of all trades, master of an unknown,” is how Dr. Dan Tawfik of the Biological Chemistry Department describes a multi-purpose enzyme, nicknamed PON. The enzyme performs a variety of jobs in the body, including breaking down harmful chemicals found in pesticides and nerve gases and ridding the arteries of plaque-forming clumps of LDL (“bad cholesterol”) that can lead to arteriosclerosis. But these are just sidelines for PON, whose original function remains shrouded in its evolutionary past. Not only does it “moonlight,” accomplishing various jobs on the side, but it performs them inconsistently in different people, doing a sloppy job for some, a more efficient one for others. This is due to variations in its encoding gene. Referred to as polymorphism, this multiplicity of forms means, for instance, that some people’s PONs are up to 50 times more active than those of others.
How does one enzyme display such diverse performances? Very little was known about its workings when Tawfik and postdoctoral fellow Dr. Amir Aharoni decided to take a closer look.
To examine protein function, scientists apply crystallization, a process that yields X-ray images that are then translated into 3-D models. But crystallizing PON enzymes filtered from the blood - a process that yields only minuscule amounts - had proven difficult. So the team set out to create pure, genetically engineered proteins, generated in large enough quantities to crystallize and analyze them in detail.
The method they used, known as “directed evolution,” works on two principles of natural, Darwinian evolution: genetic diversity and survival of the fittest. Genetic diversity was created in the lab by inducing random mutations in the gene encoding the PON enzyme. The mutated enzymes were then inserted into bacterial cells, which were allowed to grow and multiply. Selection took place in the lab, with the scientists substituting for nature, deciding which were “fit” according to certain criteria, recombining them with other mutations and selecting again. Eventually, they found several versions of the enzyme that proved ideal for undergoing the procedures needed to solve its structure.
Studying the bacterial forms of the protein, the team succeeded in revealing many aspects of its function, including how its various forms in people differ in their ability to perform assigned tasks. They discovered, for instance, that the enzyme is shaped something like a six-bladed propeller and that modifications in its structure tend to take place near its active site (which performs the actual work of the enzyme) in a way that causes instability in the scaffold of the enzyme. They also got a good glimpse of how PON “roosts” on the HDL (“good cholesterol”) and the role it may play in sucking up oxidized lipids from the LDL (“bad cholesterol”) in arteries.
Once the main structure was solved, the research team tackled a new goal: that of creating new PON variants that would be even better at specific tasks than naturally occurring ones. Again using directed evolution, they came up with enzymes that could specialize in the chemical clean-up of harmful pesticides or reduce potential risk factors in heart disease. Interestingly, as the enzyme specialized in one task, it lost its ability to do others. “It’s similar to the jack-of-all-trades who takes up carpentry. Eventually he forgets how to do the plumbing and tile-laying,” says Tawfik.
Now that the team has a better picture of how differences in PON’s structure affect its actions in the test tube, they plan to focus on how it works inside the body. Research in this field might advance the treatment of arterio-sclerosis as well as of neuronal damage arising from exposure to harmful chemicals.
Other scientists collaborating in this research are Leonid Gaidukov of the Biological Chemistry Department; Prof. Israel Silman and Lilly Toker of the Neurobiology Department and Prof. Joel Sussman and Dr. Michal Harel of the Structural Biology Department.
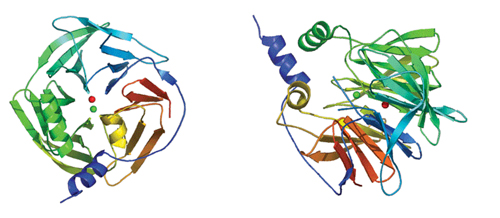
To the uninitiated, diagrams of protein structures may look like colorful tangles of ribbon or free-style children’s art projects. Yet in proteins, as in architecture, form follows function.
Each protein consists of amino acids linked together in a long chain that folds and twists into the highly intricate shape in which it performs its specialized function.
When Sir John Kendrew and Max Perutz were awarded the Nobel Prize in 1962 for solving the first three-dimensional structures of proteins, they had spent over twenty years on the task. Even with today’s improved methods, much trial-and-error work is still involved, and scientists may spend months, even years, solving a protein’s structure.
The Israel Structural Pro-teomics Center (ISPC), based at the Weizmann Institute and headed by Prof. Joel Sussman of the Structural Biology Depart-ment, has made streamlining this process its main goal. Having tackled up to 30 proteins at a time in its pilot year, the Center intends to raise that number to several hundred in the coming years. Supported by the Ministry of Science, Culture and Sports, the ISPC facility is open to scientists throughout Israeli academia and industry interested in solving protein structures.
A major partner in a network of European structural proteomics centers (which Weizmann scientists were influential in establishing), the ISPC aspires to lead the way in protein research and open up new avenues in drug design and disease treatments.
Dr. Tawfik’s research is supported by the Y. Leon Benoziyo Institute for Molecular Medicine; the Dolfi and Lola Ebner Center for Biomedical Research; the Estelle Funk Foundation; the Dr. Ernst Nathan Fund for Biomedical Research; the Henry S. and Anne Reich Family Foundation; the Harry and Jeanette Weinberg Fund for Molecular Genetics of Cancer and the Eugene and Delores Zemsky Charitable Foundation, Inc. He is the incumbent of the Elaine Blond Career Development Chair.