Despite their bad rap, bacteria can actually be quite helpful. In fact, there are about 10 times as many bacteria in the human gut as there are cells in the body, and evidence is mounting that these play a very important role in maintaining our health: They manufacture vitamins, aid in food digestion and “train” our immune system. The latter may be crucial for preventing allergies and autoimmune disease.
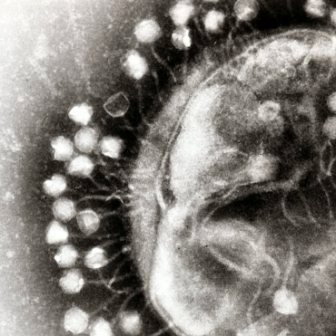
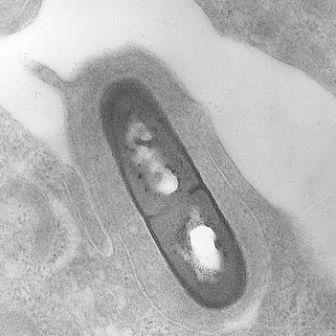
Outside the human intestine, bacteria are known to be attacked by viruses called phages, and these can sometimes kill large numbers of bacteria in a short time. The question is: Are the gut bacteria also susceptible to phage infection? And, if so, what viruses are involved and which bacteria do they attack? These questions have not received much attention until now; they were the focus of recent research by
Dr. Rotem Sorek of the Molecular Genetics Department.
The findings recently appeared in
Genome Research.
To investigate the viral population of the gut, the researchers had use of a surprising “catalog”: pieces of viral DNA that are stored in the immune systems of bacteria. The apparatus, called CRISPR, defends bacteria against viruses. When a virus gets inside a bacterial cell, CRISPR steals a small bit of the virus’s DNA and stores it in the part of its own genome that is designed specifically for this immune function. The next time the same virus infects the cell, its immune system will use the stored viral DNA to identify and destroy the invader, something like the way that antibodies work in the human body. For the scientists, these sequences lined up in the bacterial immune systems could be read as a sort of historical record of the viruses that have attacked the various gut bacteria.
The research team, including then postdoctoral fellow Dr. Adi Stern and undergraduate student Eran Mick, began with a large database of the DNA sequences of gut bacteria taken from 124 people in Europe. After developing a computational method that helped them identify the immune segments in the bacterial genome, the scientists found more than 50,000 of these immune sequences. On the basis of these sequences, they were able to identify hundreds of previously unknown viruses that infect gut bacteria.
Besides giving researchers the largest collection of information to date on phages in the human gut, the team’s findings have yielded new insights on the relationships between gut bacteria, the viruses that infect them and the humans who host them. For example, the scientists discovered that large groups of people share the same viruses, and about 80% of the viruses the team identified were found in more than one person. A comparison with samples from Americans and Japanese showed that they, too, shared the same strains. Considering the wide variety of viruses generally found in nature, this is a surprising finding; the scientists believe it may be tied to fact that the gut is a closed environment.
Another finding was that viruses are sometimes inserted into the bacterial DNA in their entirety. Sorek: “Phages occasionally contribute to the bacteria genes for antibiotic resistance. In return, the bacteria host the phages’ DNA and pass it from person to person. It’s a tradeoff that has evolved to benefit both sides.”
A comprehensive database of phages and the gut bacteria they infect may have implications for research on human health. For instance, if a certain bacterium is known to provide protection against allergies, one could check the effects of the virus that attacks that bacterium. “The ultimate goal,” says Sorek, “would be to create a vaccine against that virus, which would help the bacterium. In other words, aiding the bacterial immune system could indirectly boost the human immune system.”
The good, the bad and the bacterium
In biology, as in archaeology, digging down from one layer to the next can completely change our view of reality. Thus a move from one layer of cellular regulation – DNA – to the next – RNA – revealed a whole array of differences between two related strains of bacteria that had previously been hidden. In addition, the comparison of RNA sequences in the two strains of
Listeria – one of which causes a type of food poisoning and the other is harmless – uncovered a previously unknown mechanism for regulating the activation of
Listeria genes.
The findings appeared in
Molecular Systems Biology.
Dr. Rotem Sorek and his group – Omri Wurtzel, Dr. Iris Karunker-Hazan and Sarit Edelheit – used advance sequencing techniques to map all of the RNA for the two strains, but they actually focused on the sequences that don’t encode proteins. Though the two strains are highly similar, genetically, the team discovered a group of about 80 RNA sequences that exist in the disease-causing version but not in its “peace-loving” cousin. Now the researchers plan to check whether these sequences are responsible for the aggressiveness of the first strain.
In addition, the scientists discovered a unique pairing system in which each member of an RNA pair puts the brakes on the other. The researchers think that this system, which regulates both the activation and the silencing of genes in one mechanism, may be common in bacteria.
Dr. Rotem Sorek's research is supported by the Y. Leon Benoziyo Institute for Molecular Medicine; the Abisch Frenkel Foundation for the Promotion of Life Sciences; the Phyllis and Joseph Gurwin Fund for Scientific Advancement; the Leona M. and Harry B. Helmsley Charitable Trust; the Robert Rees Applied Research Fund; and the Alhadeff Research Award. Dr. Sorek is the incumbent of the Rowland and Sylvia Schaefer Career Development Chair in Perpetuity.