Are you a journalist? Please sign up here for our press releases
Subscribe to our monthly newsletter:
The idea of proteins that can be designed on computers for specific functions has been a cutting-edge concept that has stubbornly remained “in the future.” New research at the Weizmann Institute of Science may bring that future a bit closer. By going back to nature’s drawing board – evolution – the scientists have created new proteins based on “existing natural parts,” that carry out their intended function with flying colors. This research was reported yesterday in the Proceedings of the National Academy of Science, PNAS.
The research was conducted by Dr. Sarel Fleishman, research students Dror Baran, Maria Gabriele Pszolla, Gideon Lapidoth, and other members of Fleishman’s group in the Weizmann Institute of Science’s Biomolecular Sciences Department. In Sarel’s lab, proteins are designed with computer-based programs that enable them to generate new structures – for example, antibodies or enzymes – that do not exist in nature. If they want a protein that will perform a specific action, say bind to another protein or carry out a chemical reaction, they can compute, from beginning to end, the genetic sequence that will line up amino acids in the proper order and cause the protein to fold into the correct three-dimensional shape. Such proteins could, in theory, usher in a new age of custom-designed drugs and catalysts, but the challenges of planning of large biological molecules are immense.
They created a sort of symbiotic evolution – the design programs evolving along with the experimental tests
The team then asked a simple question: “What does a natural enzyme or antibody have that the artificial proteins don’t? Why are two structures with similar makeup so different when it comes to the way they perform inside a biological system?”
The group focused their attention on some parts of natural antibodies or enzymes that don’t make it into the computer designs which start from scratch – particularly structures called “loops,” which are inherently unstable and “non-ideal” and therefore challenging when it comes to computational prediction. These “non-ideal” loops can be often be found at the very center of the active regions – those that recognize a target or bind to or cleave another molecule.
To incorporate these parts, the researchers decided to design a functioning antibody from existing parts, rather than building one from scratch. They broke the structures found in natural antibodies down into segments, including the loops and other supporting features. In effect, the researchers tinkered with ready-made parts, similar to the way evolution works.
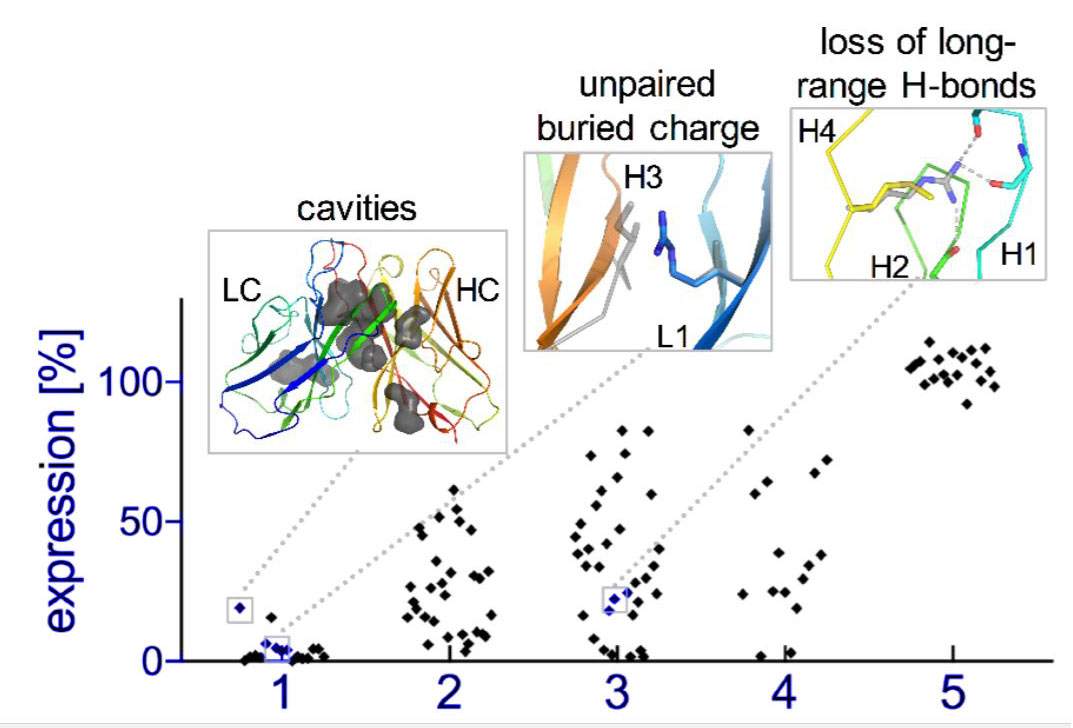
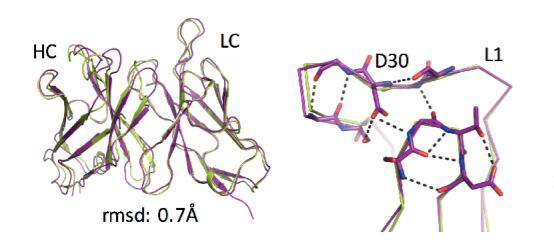
Natural evolution is, of course, a very slow process: A new family of antibodies can be tens of millions of years in the making. So the researchers went back to the computerized planning process, this time armed with their new insight. The new designs were then tested experimentally in the lab, a few dozen new antibodies at a time. Initially, the designs performed poorly, but through five design-build-test cycles, the researchers uncovered some general rules for designing antibodies. In essence, they created a sort of symbiotic evolution – the design programs evolving along with the experimental tests, each pushing the other forward. To demonstrate the feasibility of this concept, the team created artificial antibodies that targeted insulin, and they characterized these molecules down to the resolution of single atoms.
In future experiments, the scientists plan to design artificial antibodies modeled on those of camels and llamas. Why these particular animals? If a human antibody or one from any number of common animals has 200 amino acids, in camels and llamas they are made of just 100, and yet are stable and effective. This could make the design and production of artificial antibodies for human conditions more efficient, and might have relevance for designing new diagnostics and therapeutics.
Dr. Sarel-Jacob Fleishman’s research is supported by the Henri Gutwirth Fund for Research; the Rothschild Caesarea Foundation; Sam Switzer, Canada; Anne Christopoulos, Canada; and Carolyn Hewitt, Canada. Dr. Fleishman is the incumbent of the Martha S. Sagon Career Development Chair.