A synthetic device made of biological molecules can be programmed to search for and identify exact DNA sequences inside living cells – and reject any of those sequences that contain errors, however tiny. The Weizmann Institute researchers who invented the device believe that the concept behind it may lead to the development of new, highly sensitive diagnostic techniques, as well as enhanced methods for creating interfaces between natural and synthetic biological molecules.
Dr. Tuval Ben Yehezkel and Tamir Biezuner in the lab of
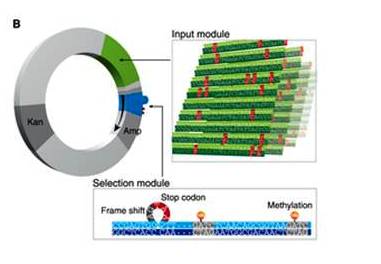
Prof. Ehud Shapiro of the Biological Chemistry, and Computer Science and Applied Mathematics Departments created numerous copies of the DNA-based devices – each containing an identical, preprogrammed genetic sequence hooked up to a different target gene – and inserted them into bacterial cells. Inside the cells, the synthetic infiltrators, like tiny moles, recruited the cell’s “proofreaders”: internal repair mechanisms that normally, during cell replication, check for mismatches between the genetic “letters” A,T,G and C in a new DNA strand and those in the parallel sequence of the double strand. Normally, these proofreaders would snip out any offending letters on one strand and call on the paired strand of DNA to try substituting the information. The synthetic device, however, picked up on the error-correction activity and co-opted this mechanism, using it instead to reprogram itself to destroy the cell. Thus only cells containing the complete, correct sequence remained at the end of the process.
In the experiment, the devices were programmed to preserve a DNA sequence that does not confer any selective advantage in E. coli cells. This ability to work with any DNA segment, regardless of its use in the cell, highlights its advantage over other methods: The other methods often rely on a gene’s functionality to select for it using classical Darwinian selection, or are liable to miss tiny errors in the sequence. In the future, slightly more sophisticated versions of the device could be used by researchers working with artificial genetic sequences to ensure their accuracy. The researchers think that future devices based on their concept could be employed in medicine to seek out hard-to-detect genes – for example harmful mutations in fetal cells in the mother’s blood or cancer-causing mutations that appear in just a few cells.
The findings, which appeared recently in
PLoS One, incorporate aspects of two fairly new fields of research: synthetic biology and biological computing. They have demonstrated how a man-made genetic sequence can be inserted into a living cell and interface there with the cell’s natural mechanisms – a feat that has been a major, ongoing challenge for synthetic biologists. And with the help of those mechanisms, this synthetic construct performs like a simple computing device, in which DNA inputs – sequences that may or may not contain errors – are processed to arrive at an output – in this case the preservation or elimination of those sequences. Shapiro says: “Future work in this direction may bring about the integration of synthetic DNA devices into ever more complex cellular environments. This, in turn, could lead to a wide variety of applications.”
Prof. Ehud Shapiro’s research is supported by the Paul Sparr Foundation; and the European Research Council. Prof. Shapiro is the incumbent of the Harry Weinrebe Professorial Chair of Computer Science and Biology.