In recent years, a number of controversial claims have been made about the female mammal’s egg supply – that it is renewed over her adult lifetime (as opposed to the conventional understanding that she is born with all of her eggs), and that the source of these eggs is stem cells that originate in the bone marrow. Now, Weizmann Institute scientists have disproved one of those claims and pointed in new directions toward resolving the other. Their findings, based on an original method for reconstructing lineage trees for cells, were published in the online journal PLoS Genetics.
The method, developed over several years in the lab of
Prof. Ehud Shapiro of the Institute’s Biological Chemistry, and Computer Science and Applied Mathematics Departments, uses mutations in specific genetic markers to determine which cells are most closely related and how far back they share a common parent cell, to create a sort of family tree for cells. Shapiro and members of his lab, including Drs. Shalev Itzkovitz and Rivka Adar, together with
Prof. Nava Dekel and research student Yitzhak Reizel of the Biological Regulation Department, used their method to see if ova could be descended from bone-marrow stem cells. Their findings indicated that any relationship between the two types was too distant for one to be an ancestor of the other.
These scientists also found, surprisingly, that the ova of older mice had undergone more cell divisions than those of younger mice. This could be the result of replenishment during adulthood; but an alternate theory holds that all eggs are created before birth, and those that undergo fewer divisions are simply selected earlier on for ovulation. Further experimentation, says Shapiro, will resolve the issue.
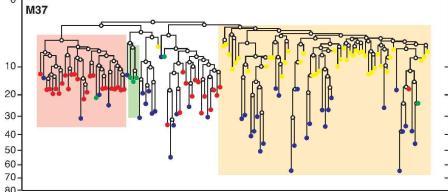
Cell lineage trees are similar to modern evolutionary and taxonomic trees based on genome comparisons among organisms. Shapiro and his team used mutations in cells that are passed on to daughter cells over an organism’s lifetime (though not on to the next generation). By comparing a number of genetic sequences called microsatellites – areas where mutations occur like clockwork – they can place cells on trees to reveal their developmental history.
A number of papers published by Shapiro, his team and collaborators in recent months have demonstrated the power and versatility of this method. One study, for instance, lent support to the notion that the adult stem cells residing in tiny crypts in the lining of the colon do not harbor, as thought, “immortal DNA strands.” Immortal strands may be retained by dividing stem cells if they always relegate the newly synthesized DNA to the differentiating daughter cell and keep the original strand in the one that remains a stem cell.
A second study addressed an open question about developing muscle cells. Here they found that two kinds of progenitor cell – myogenic cells, which eventually give rise to muscle fiber, and non-myogenic cells – found within the same muscle are more closely related than similar cells in different muscles.
One immediate advantage of the cell lineage analysis method developed by Shapiro’s team is that it is non-invasive and retrospective, and as such can be applied to the study of human cell lineages. Most other studies of development rely on genetically engineered lab animals in which the stem cells are tagged with fluorescent markers. In addition to providing a powerful new research method that does not rely on such markers, Shapiro believes that it could one day be adapted as a diagnostic tool that might, for instance, reveal the history of an individual’s cancer and help doctors determine the best course of treatment.
Also participating in this research were Noa Chapal-Ilani, Adrian Jinich and Drs. Elad Segev, Eran Segal and Yosef Maruvka, of the Computer Science and Applied Mathematics Department; Zipora Marx, Inna Horovitz and Adam Wasserstrom of the Biological Chemistry Department; Drs. Judith Elbaz and Nava Nevo of the Biological Regulation Department; Dr. Avi Mayo of the Molecular Cell Biology Department; Drs. Gabi Shefer and Irena Shur, and Prof. Dafna Benayahu of Tel Aviv University; and Prof. Karl Skorecki of the Technion and Rambam Medical Center, Haifa.
Prof. Nava Dekel's research is supported by the Jeanne and Joseph Nissim Foundation for Life Sciences Research; the M.D. Moross Institute for Cancer Research; the Y. Leon Benoziyo Institute for Molecular Medicine; the Yeda-Sela Center for Basic Research; the Willner Family Center for Vascular Biology, which she heads; the Dwek Family Biomedical Research Fund; the J&R Foundation; the estate of John Hunter; and the Allyson Kaye Charitable Trust. Prof. Dekel is the incumbent of the Philip M. Klutznick Professorial Chair of Developmental Biology.
Dr. Eran Segal's research is supported by the Cecil and Hilda Lewis Charitable Trust; the Carolito Stiftung; the Kahn Family Research Center for Systems Biology of the Human Cell; and the European Research Council.
Prof. Ehud Shapiro's research is supported by the Paul Sparr Foundation; Miel de Botton, UK; the Carolito Stiftung; and the European Research Council. Prof. Shapiro is the incumbent of the Harry Weinrebe Professorial Chair of Computer Science and Biology.