One person’s trash, they say, is another’s treasure. For Amit Zeisel, a research student in the group of
Prof. Eytan Domany of the Physics of Complex Systems Department, and Dr. Wolfgang Köstler, an M.D. conducting Ph.D. research in the lab of
Prof. Yosef Yarden of the Biological Regulation Department, reluctance to throw out seemingly useless information led to a revision in our understanding of how certain RNAs are produced in the cell.
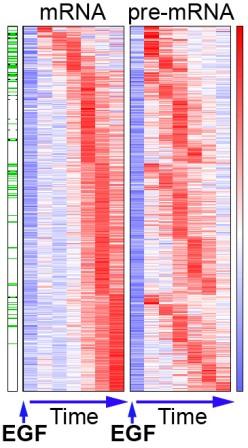
Ongoing collaboration between the labs of Domany and Yarden has been revealing the timing of the production of messenger RNA (mRNA), the strands that carry the protein-making instructions encoded in the genes out of the cell nucleus. After the cell receives a signal to activate a series of genes, the different mRNA levels begin to rise in what appear to be coordinated steps, some peaking within half an hour, others after one or two hours. These signals and the ensuing mRNA production are the first steps of nearly all cellular activity, and understanding the timing – which mRNAs are produced right away and which peak only several hours later – is helping researchers to understand some of the most basic processes in health and disease, for instance, how cancer develops.
The new finding arose out of some previous research between the labs involving apparent waste in the process of mRNA production. Starting with the pre-mRNA – an exact transcript of the gene code “written” in the DNA – short sequences called exons are snipped out and spliced together, while the pieces in between, called introns, are left out of the final mRNA strand. For their study, Zeisel and Köstler were using a research tool that enables them to track those pieces – individual exons and introns – over time.
The two were looking at variations in the exon splicing to see if certain versions might be tied to the timing of mRNA production.
Most researchers simply discount the introns; after all, it is only the exons that go into the final instruction list. Even though their study only involved exons, Zeisel and Köstler hit upon the idea that the intron information might come in handy and left some of it in the data sets. That is when the researchers noticed something surprising: The shape, height and timing of the production peaks they were seeing – which belonged to pre-mRNAs – looked quite different from those of the corresponding mRNAs.
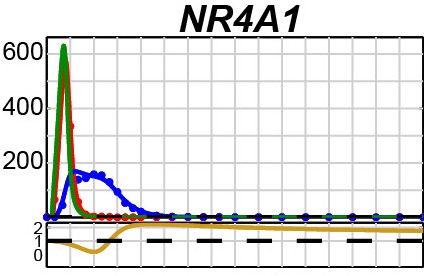
The team realized that because introns are excluded from the mRNA, the accidental discovery presented a novel way of measuring pre-mRNA dynamics on a genome-wide level. The findings implied that pre-mRNA is produced on a different schedule than mRNA and not, as scientists had assumed, at the same rate. To check this idea, they conducted a carefully designed experiment using both introns and exons. Their results showed that in a number of cases, pre-mRNA levels spiked quickly – rising to a much higher level than the mRNA peaks that appeared later on – and fell off just as quickly.
Afterward, the team took their insight to other Institute research groups to see if the same pattern is found in different cell types and in different situations. And indeed, in the immune cells researched in the group of
Prof. Steffen Jung as well as in the embryonic stem cells in the lab of Dr. Yoav Soen, certain RNAs exhibited patterns that were similar to those Yarden, Domany and their students had seen in their original experiment.
The scientists termed these early, high and narrow pre-mRNA peaks “production overshoot.” “We begin observing these cells as they receive a signal that initiates a particular chain of events,” explains Domany. If these cells suddenly need large amounts of a certain protein, a fast jumpstart in pre-mRNA production might get things moving.” He compares it to merging from a slow on-ramp into fast-moving freeway traffic: Instead of adjusting his foot on the gas to the level needed for cruising at the higher speed, a driver will put the pedal to the metal for a minute to get up to speed and then ease off again.
In addition to challenging some previous assumptions about the steps in the process that leads to protein production, Domany points out that having a second “known” about this multi-step cellular activity can enable researchers to deduce other unknowns. For instance, if one knows the temporal profiles of both pre-mRNA and mRNA, one can calculate both the pre-mRNA production rate and the mRNA degradation rate. In fact, this finding is helping to paint a detailed picture of mRNA regulation. In that picture, pre-mRNA overproduction and the carefully timed degradation of the mRNA strand work together as a sort of combination gas-and-brake mechanism, adjusting its levels over time. Now, the team is adding an investigation of protein levels to their research, in hopes of obtaining a fuller picture of the entire procedure – from the instant the pre-mRNA is copied off the gene to the rolling of the proteins off the production line. And, they say, they have learned a valuable lesson: Don’t be surprised to find paradigm-changing information sitting right under your nose – in the discard pile.
Prof. Eytan Domany’s research is supported by the Kahn Family Research Center for Systems Biology of the Human Cell, which he heads; the Mario Negri Institute for Pharmacological Research - Weizmann Institute of Science Exchange Program; the Leir Charitable Foundations; and Mordechai Segal, Israel. Prof. Domany is the incumbent of the Henry J. Leir Professorial Chair.
Prof. Yosef Yarden’s research is supported by the European Research Council; the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation; the Aharon Katzir-Katchalsky Center, which he heads; the M.D. Moross Institute for Cancer Research; the Kekst Family Institute for Medical Genetics; the Steven and Beverly Rubenstein Charitable Foundation, Inc.; Julie Charbonneau, Canada; and the Marvin Tanner Laboratory for Research on Cancer. Prof. Yarden is the incumbent of the Harold and Zelda Goldenberg Professorial Chair in Molecular Cell Biology.