Are you a journalist? Please sign up here for our press releases
Subscribe to our monthly newsletter:
All life depends on proteins; thus studies in pretty much all the life sciences – including biomedical and pharmaceutical research – require the use of proteins. But many proteins, particularly large or human ones, are unstable; they can’t be produced in the bacterial systems that have today become standard, and may require expensive and inefficient alternate systems. Instability also arises because many proteins are sensitive to heat, environmental conditions or high concentrations, or else just have a short lifespan. For scientists, the stability problem can be critical. Promising therapeutics have even been known to fail because the proteins were not sufficiently stable.
Dr. Sarel Fleishman and his research student Adi Goldenzweig of the Weizmann Institute of Science’s Biomolecular Sciences Department have now proposed a solution to this problem. In Fleishman’s lab, computational tools are used to design new protein structures. Taking on the ambitious task of improving protein stability, they succeeded beyond their dreams: The method they created can be used to design stable versions of nearly any protein. This could assist the research of many groups around the world; and it may, in the future, lead to better protein-based drugs.
The researchers began the study with a protein called acetylcholinesterase (AChE). AChE breaks down a neurotransmitter that nerves use to communicate with muscles. When AChE malfunctions, muscles − including cardiac and lung muscles − are stuck in a contracted state. A large protein made up of 550 amino acids, AChE is susceptible to falling apart when the temperature changes and its large size leaves room for mistakes in production or folding. The scientists’ plan was to introduce stabilizing mutations into the protein’s sequence where needed. A hefty protein like AChE would require dozens of mutations: The challenge was that any one mutation could potentially interfere with the protein’s function, and combining them could lead to further disruptions.

Preserving function, increasing stability
Fleishman and Goldenzweig first needed to know which parts of the protein’s sequence of amino acids are vital. They compared human AChE with hundreds of proteins that perform similar functions in animals, from fish to humans, on the assumption that if an amino acid was conserved throughout the protein’s natural evolution, it was probably necessary. The next step – introducing the stabilizing mutations – called for special software, Rosetta, which has been developed in collaboration between dozens of labs all over the world, including Fleishman’s.
Fleishman explains: “We look at the forces that govern protein stability − for example, the electrostatic force that attracts a negative charge to a positive one. The attraction and repulsion between the charges on amino acids give the protein its shape and determine how well it will withstand higher temperatures. Thus we look for amino acid substitutions that change the nature of the attraction (or repulsion) and, like tightening dozens of screws in the structure, stabilize the protein.”
Goldenzweig, a self-confessed technophobe, learned the Rosetta software as well as the programming language Python and used them to write algorithms for designing a completely revamped protein. The predicted stable version of AChE ultimately entailed the introduction of 51 different mutations: that is, almost 10% of the amino acids in the sequence − an unprecedented number for design. Moshe Goldsmith from Tawfik’s lab then tested the predicted version experimentally. The protein showed dramatic improvements: It can be produced in substantial amounts in bacterial systems; and it is functional at temperatures of up to 70˚C, while the original protein breaks down at around 50˚C.
“With such a large number of mutations, there should have been almost no chance that the function of the protein would not be affected,” says Fleishman. “When we sent our best design to Prof. Tawfik,” says Goldenzweig, “we were sure we would, at the very least, have to go back and make further adjustments.” But when he had the experimental results, Tawfik emailed to them: “Brilliant!”
The Weizmann Institute’s Israel Structural Proteomics Center then produced large amounts of the designed protein and determined its 3D structure to atomic resolution; despite the 51 mutations, the protein’s active site was virtually indistinguishable from that of the human protein.
The pros of PROSS
Fleishman and Goldenzweig realized that they had figured out how to stabilize not only one protein, the computer-design method they had developed suggested a basic plan for reinforcing the “body” of nearly any protein without touching its moving parts. Together with Dr. Jaime Prilusky, the researchers created a webserver, which they named the Protein Repair One Stop Shop (PROSS). Goldenzweig says: “We expect researchers to apply PROSS to their proteins just once; they should then be so stable, we won’t get return customers.”
Fleishman, knowing that many researchers struggle with stability issues, began to describe the algorithm in talks and conferences. Researchers from Georgia Tech who had heard about it, then applied the PROSS method, using an automatic query to the webserver. Fleishman and his lab were not even aware of the new users until they received an email from the Georgia Tech team with promising results on a stabilized glaucoma protein.
Other users also reported success, verifying the method. Fleishman: “Anyone can use it. You put your protein in, and a few hours later it gives you just a handful of versions predicted to be stabilizing, which can then be easily tested in the lab − rather than the hundreds of thousands of options you would normally get by trying to just combine mutations from the databases.” “Even before the paper was published, news of the PROSS webserver began to spread, and we have already had more than one hundred requests to use it,” says Goldenzweig.
We expect researchers to apply PROSS to their proteins just once; they should then be so stable, we won’t get return customers
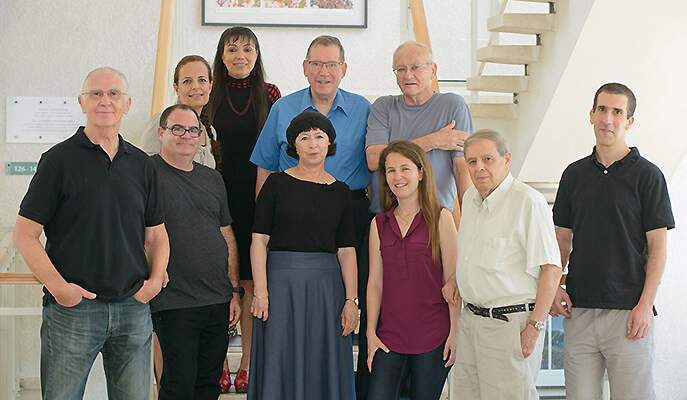
Silman, Sussman and Tawfik are all coauthors of the new paper that was recently published in Molecular Cell. This paper reports the stabilization of AChE and four other challenging proteins. Looking back, Fleishman says that the extra effort in tackling a protein that tested Goldenzweig and the rest of the team to the limits of their skills more than paid off.
Fleishman and his group already have a list of proteins they plan to improve in the future. He is particularly optimistic about the use of this method for the development of new protein-based drugs. Such proteins might be produced in bacteria or other inexpensive cellular systems; and the yield would be improved, as the proteins would be resistant to misfolding and less responsive to adverse environmental conditions. In addition, they might be designed to remain stable in warm environments, or for long periods, thus, for example, helping get vaccines to people who don’t have refrigeration, or effective antidotes such as AChE for nerve-gas poisoning to those who are not near a hospital.
Dr. Sarel-Jacob Fleishman's research is supported by the Rothschild Caesarea Foundation; Sam Switzer, Canada; and the European Research Council. Dr. Fleishman is the incumbent of the Martha S. Sagon Career Development Chair.